Im trying to segment 3D volumes using a 3D uNet network. Ive reached a stage where I am getting very good validation loss using CrossEntropy and BCE
idx: 0 of 53 - Validation Loss: 0.029650183394551277
idx: 5 of 53 - Validation Loss: 0.009899887256324291
idx: 10 of 53 - Validation Loss: 0.05049080401659012
idx: 15 of 53 - Validation Loss: 0.02019292116165161
idx: 20 of 53 - Validation Loss: 0.04724293574690819
idx: 25 of 53 - Validation Loss: 0.02810296043753624
idx: 30 of 53 - Validation Loss: 0.02642594277858734
idx: 35 of 53 - Validation Loss: 0.029894422739744186
idx: 40 of 53 - Validation Loss: 0.04158024489879608
idx: 45 of 53 - Validation Loss: 0.04574814811348915
idx: 50 of 53 - Validation Loss: 0.05406259000301361
I assumed my network is performing very well so i wrote a script to visualize my network outputs against their respective targets. What I get is something very different, not something that justifies this loss.
The samples are of depth 32 and I’ve outputted each z-plane as a single image.
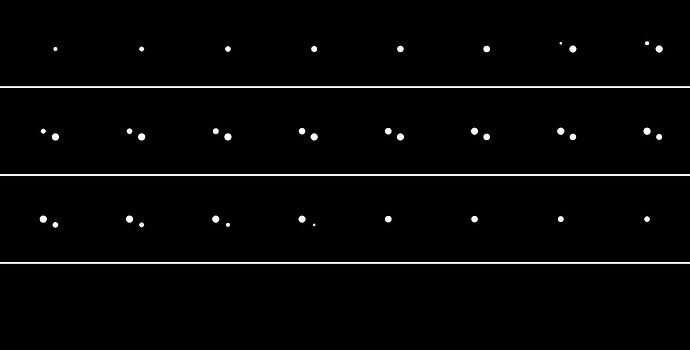
Here is the target:
And the predicted output:
All samples are like this, not one of them accurately represent the target with the reported loss… so I ask is my loss wrong? What should I look into to fix this?
Thanks