Hi,
I am fairly new to torch and trying to use LSTM to predict time series data. I pretty much followed this tutorial, except I changed LSTMCell to LSTM. Here is the code I end up with:
========== IMPORTS ==========
import pandas as pd
import torch
import pylab
import numpy as np
import torch.nn as nn========== PARAMETERS ==========
data_file = “…/data/ecg/qtdbsel102.txt”
input_dim = 1
hidden_dim = 51
batch_size = 100
output_dim = 1
num_layers = 2
learning_rate = 0.1
num_epochs = 100
use_gpu = True========== READ ECG DATA ==========
df = pd.read_csv(data_file,sep=“\t”,header=None,names=[“t”,“v1”,“v2”],dtype={“t”:float,“v1”:float,“v2”:float})
x =
y =
for i1 in range(len(df)//batch_size-1):
x.append(list(df[“v1”].iloc[i1*batch_size:(i1+1)*batch_size]))
y.append(list(df[“v1”].iloc[(i1*batch_size+1):((i1+1)*batch_size+1)]))
x = torch.from_numpy(np.array(x)).float()
if(use_gpu==True):
x = x.cuda()
y = torch.from_numpy(np.array(y)).float()
if(use_gpu==True):
y = y.cuda()========== CREATE MODEL ==========
class LSTM(nn.Module):
def __init__(self, input_dim, hidden_dim, batch_size, output_dim=1, num_layers=2): super(LSTM, self).__init__() self.input_dim = input_dim self.hidden_dim = hidden_dim self.batch_size = batch_size self.num_layers = num_layers # Define the LSTM layer self.lstm = nn.LSTM(self.input_dim, self.hidden_dim, self.num_layers) # Define the output layer self.linear = nn.Linear(self.hidden_dim, output_dim) def init_hidden(self): # This is what we'll initialise our hidden state as return (torch.zeros(self.num_layers, self.batch_size, self.hidden_dim), torch.zeros(self.num_layers, self.batch_size, self.hidden_dim)) def forward(self, input): # Forward pass through LSTM layer # shape of lstm_out: [input_size, batch_size, hidden_dim] # shape of self.hidden: (a, b), where a and b both # have shape (num_layers, batch_size, hidden_dim). lstm_out, self.hidden = self.lstm(input.view(len(input), self.batch_size, -1)) # Only take the output from the final timetep # Can pass on the entirety of lstm_out to the next layer if it is a seq2seq prediction y_pred = self.linear(lstm_out) return y_pred.squeeze(2)model = LSTM(input_dim,hidden_dim,batch_size,output_dim,num_layers)
if(use_gpu==True):
model.cuda()
criterion = torch.nn.MSELoss(reduction=“sum”)
optimizer = torch.optim.LBFGS(model.parameters(), lr=learning_rate)========== TRAIN MODEL ==========
loss_hist =
for i in range(num_epochs):
# step through
print('STEP: ', i)
def closure():
optimizer.zero_grad()
y_pred = model(x)
loss = criterion(y_pred, y)
print(‘loss:’, loss.item())
loss.backward()
return loss
optimizer.step(closure)
# plot progress
y_pred = model(x)
loss_hist.append(criterion(y_pred,y).item())
pylab.figure()
pylab.plot(y.cpu()[0].numpy(),“b”)
pylab.plot(y_pred.cpu().detach()[0].numpy(),“r”)
pylab.savefig(“%05d.jpg”%(i))plot loss history
pylab.figure()
pylab.plot(loss_hist)
pylab.savefig(“loss_hist.jpg”)
pylab.close()
The data file is from here.
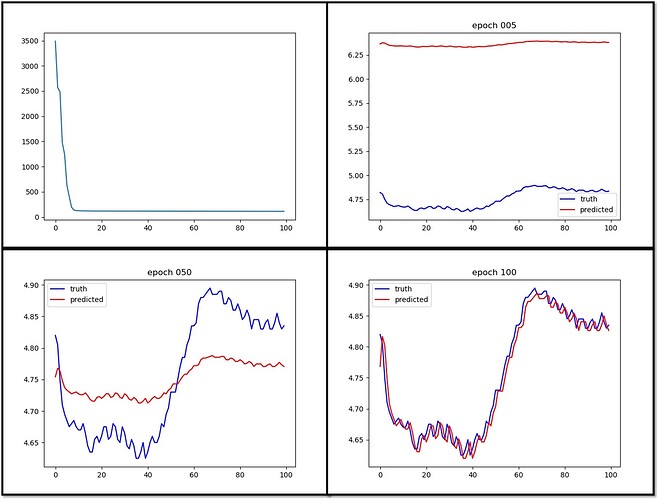
The first plot is the loss history.
I currently have problems.
- It seems prediction gets look similar to the truth very quickly but it takes a very long training time to get the amplitude aligns well with the truth. This is counter intuitive to me because the large scale amplitude should be much easier to train than the high frequency details.
-
In the trained output, there is a phase shift between the prediction and truth. Is there a way to eliminate it?
-
In general, is it valuable to feed the time axis as another input to the training?
Thanks.