Per-example and mean-gradient calculations work on the same set of inputs, so PyTorch autograd already gets you 90% of the way there. For an illustration: consider the problem of backprop in a simple feedforward architecture.
Each layer has a Jacobian matrix and we get the derivative by multiplying them out.

While any order of matrix multiplication is valid, reverse differentiation specializes to multiplying them “left to right”.
For “left-to-right” order, each op implements “vector-Jacobian” product, called “backward” in PyTorch, “grad” in TensorFlow and “Lop” in Theano. This is done without forming Jacobian matrix explicitly, necessary for large scale applications.
When dealing with non-linear functions, each example in a batch corresponds to a different Jacobian, so our backward functions do this in a batch fashion.
Autograd engine traverses the graph and feeds a batch of these vectors (backprops) into each op to obtain backprops for the downstream op.
To see how per-example calculation works, note that for matmul, parameter gradient is equivalent to computing a batch of outer products of matching activation/backprop vector pairs, then summing over the batch dimension. Activations are values fed into the layer during forward pass.
We get the per-example gradients by dropping the sum
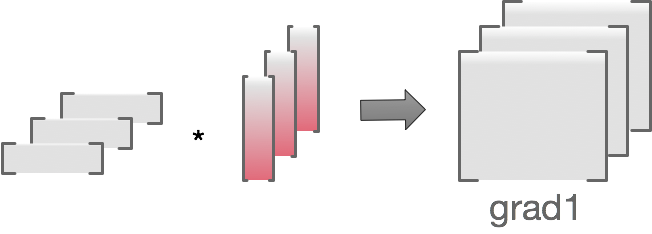
Basically it’s a matter of replacing
grad=torch.einsum('ni,nj->ij', backprops, activations)
with
grad1=torch.einsum('ni,nj->nij', backprops, activations)
Because grad1 calculation doesn’t affect downstream tasks, you only need to implement this for layers that have parameters. The autograd-hacks lib does this for Conv2d and Linear. To extend to a new layer, you would look for the “leaf” ops, look at their “backwards” implementation and figure out how to drop the “sum over batch” part.




