Hi, I am new at Pytorch and I am trying to classify Ocular Dataset with both NN and CNN. However, although my loss is decreasing through epoch, my accuracy stays same, which is because my model always predicts class zero. I have searched for a reason but nothing fixed my issue. I already have gray scaled and normalized and flatten the dataset, I shuffled before splitting the dataset into train, validation and test. I also shuffled while loadşng the data. And lastly, I tried to change batch size, learning rate and epoch size but it did not work. Can you please help me to find the problem?
I use BCEWithLegitsLoss for my loss function and SGD for optimizer.
The code is below for getting dataset:
def get_dataset(root):
#os.mkdir('/content/gdrive/My Drive/Colab Notebooks/images/1')
#os.mkdir('/content/gdrive/My Drive/Colab Notebooks/images/0')
label_data = pd.read_excel('/content/gdrive/My Drive/Colab Notebooks/labels.xlsx')
labels = label_data.iloc[:,1]
ids = label_data.iloc[:,0]
#idx = 0
#for id in ids:
#os.rename('/content/gdrive/My Drive/Colab Notebooks/images/'+ str(id) +'.jpg', '/content/gdrive/My Drive/Colab Notebooks/images/'+ str(labels[idx]) +'/'+ str(id) + '.jpg')
transform_data = T.Compose([T.Grayscale(), T.ToTensor() ])
train_dataset = ImageFolder(root=root,transform=transform_data)
images = []
labels = []
#check = 0
for image, label in iter(train_dataset):
#print(" " + str(check), end='')
T.Normalize(2, 0.5)(image)
images.append(image.view(32768))
labels.append(label)
labels = torch.tensor(labels)
images = torch.stack(images)
print("Before shuffle")
#Shuffle data
l = list(range(len(ids)))
random.shuffle(l)
train_shuffled = l[0:images.shape[0] * 5 // 7]
val_shuffled = l[len(train_shuffled):len(train_shuffled) + images.shape[0] // 7]
test_shuffled = l[len(train_shuffled) + images.shape[0] // 7:len(train_shuffled) + images.shape[0] // 7
+len(train_shuffled) + images.shape[0] // 7]
# Construct training, validation and test sets
train_data = images[train_shuffled, ...]
train_label = labels[train_shuffled]
val_data = images[val_shuffled, ...]
val_label = labels[val_shuffled]
test_data = images[test_shuffled, ...]
test_label = labels[test_shuffled]
train_dataset = OcularDataset(data = train_data, labels = train_label)
val_dataset = OcularDataset(data = val_data, labels = val_label)
test_dataset = OcularDataset(data = test_data, labels = test_label)
return train_dataset, val_dataset, test_dataset
And this is my main, train and test functions:
max_epoch = 50
train_batch = 32
test_batch = 32
learning_rate = 0.0001
device = torch.device(‘cuda’)
if(torch.cuda.is_available()):
print(“Using GPU”)
def main(): # you are free to change parameters
if(torch.cuda.is_available()):
print(“Using GPU”)
# Create train dataset loader
train_loader = DataLoader(train_set, batch_size=train_batch, shuffle=True)
# Create validation dataset loader
valid_loader = DataLoader(val_set, batch_size=test_batch, shuffle = True)
# Create test dataset loader
test_loader = DataLoader(test_set, batch_size=test_batch, shuffle = True)
# initialize your GENet neural network
model = FNet().to(device)
# define your loss function
loss_func = nn.BCEWithLogitsLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate, momentum=0.9, weight_decay=5e-04) # you can play with momentum and weight_decay parameters as wel
train_acc =
train_loss =
valid_acc =
valid_loss =
min_loss = 10000
# start training
# for each epoch calculate validation performance
# save best model according to validation performance
for epoch in range(max_epoch):
train(epoch, model, loss_func, optimizer, train_loader, train_acc, train_loss)
acc,loss = test(model, valid_loader, loss_func)
print()
valid_acc.append(acc)
valid_loss.append(loss)
if loss < min_loss:
min_loss = loss
torch.save(model, ‘best_path.pth’)
return (train_acc, train_loss, valid_acc, valid_loss)
‘’’ Train your network for a one epoch ‘’’
def train(epoch, model, loss_func, optimizer, loader,train_acc,train_loss): # you are free to change parameters
model.train()
loss_tot = 0
acc_tot = 0
prediction_tot = 0
for batch_idx, (data, labels) in enumerate(loader):
start = time.time()
# TODO:
# Implement training code for a one iteration
data = data.to(device=device, dtype=torch.float32)
labels = labels.to(device=device, dtype=torch.float32)
optimizer.zero_grad()
output = model(data)
loss = loss_func(output, labels.float().unsqueeze(1))
loss.backward()
optimizer.step()
end = time.time()
predictions = output.argmax(dim=1)
corrects = 0
for idx in range(labels.shape[0]):
if predictions[idx] == labels[idx]:
corrects += 1
#corrects = (predictions == labels).sum()
acc = corrects / labels.shape[0]
prediction_tot += labels.shape[0]
acc_tot += acc
loss_tot += loss.item()
train_acc.append(acc)
train_loss.append(loss)
print(‘Epoch: [{0}][{1}/{2}]\t’
‘Time {batch_time:.3f}\t’
‘Loss {loss:.4f} ({loss_avg:.4f})\t’
‘Accu {acc:.4f} ({acc_avg:.4f})\t’.format(
epoch + 1, batch_idx + 1, len(loader),
batch_time=end - start,
loss=loss.item(), loss_avg = loss_tot / (batch_idx+1),
acc = acc, acc_avg = acc_tot/(batch_idx+1)))
‘’’ Test&Validate your network ‘’’
def test(model, loader, loss_func): # you are free to change parameters
model.eval()
corrects = 0
samples = 0
with torch.no_grad():
for batch_idx, (data, labels) in enumerate(loader):
# TODO:
# Implement test code
start = time.time()
data = data.to(device=device) # move to device, e.g. GPU
labels = labels.to(device=device)
output = model(data)
loss = loss_func(output, labels.float().unsqueeze(1))
end = time.time()
predictions = output.argmax(dim=1)
for idx in range(labels.shape[0]):
if predictions[idx] == labels[idx]:
corrects += 1
samples += (labels.shape[0])
acc_avg = corrects/samples
print(‘Time: {batch_time:.3f}\t’
‘Loss: {loss:.4f}\t’
‘Accu {acc_avg:.4f}\t’.format(
batch_time=end-start,
loss = loss.item(),
acc_avg=acc_avg))
return acc_avg, loss
My FNet class is as follows:
Blockquote
class FNet(nn.Module):
def init(self, **kwargs):
super(FNet, self).init()
self.hidden_layer1 = nn.Linear(32768, 1024)
self.hidden_layer2 = nn.Linear(1024,256)
self.hidden_layer3 = nn.Linear(256,32)
self.output_layer = nn.Linear(32,1)
def forward(self, X):
x = F.relu(self.hidden_layer1(X))
x = F.relu(self.hidden_layer2(x))
x = F.relu(self.hidden_layer3(x))
x = self.output_layer(x)
x = torch.sigmoid(x)
return x
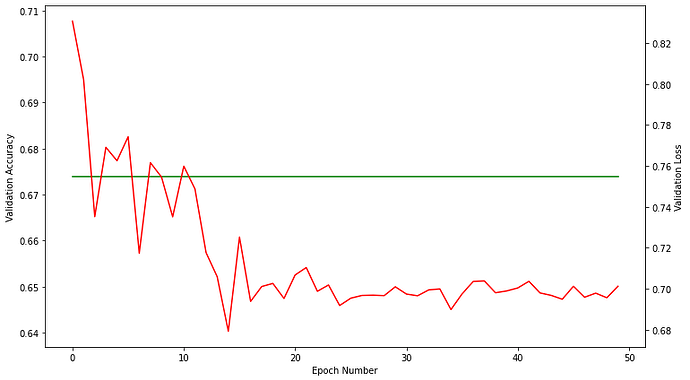
And finally, these are the plots for validation accuracy and loss in each epoch