I’m wondering if there’s a way to alter the torch.utils.data.DataLoader so that one could import FITS files into Pytorch as tensors with labels?
I’m intending to load in Astronomical images from FITS format (http://docs.astropy.org/en/stable/index.html) and presumably in the source code for DataLoader there must be some function for handling different data types such as: .jpg, .png etc.
Perhaps if there’s no simple clause for directly importing FITS files, I could be pointed towards that function and I can insert my own FITS handler clause?
Given that my code looks as follows when importing the training set:
trainset = dset.ImageFolder(root="/Documents/Image_data",transform=transform)
train_loader = torch.utils.data.DataLoader(trainset, batch_size=batch_size,shuffle=True)
…and I haven’t been able to find the data type handling functions in the source code for torch.utils.data.DataLoader or torch.utils.data.Dataset, maybe someone could point me in the right direction?
Many thanks in advance!
Update, I believe I may be able to make this work if using torchvision.datasets.DatasetFolder and setting my own loader for FITS extensions.
However when trying to use this class I get the following error:
AttributeError: module 'torchvision.datasets' has no attribute 'DatasetFolder'
Is DatasetFolder actually supported by torchvision at this point in time?
you could specify your own folder as shown in this post
I’m unsure as to where data.Dataset comes from in this class.
I’ve implemented the method you suggested above:
class FitsFolder(DatasetFolder):
EXTENSIONS = ['.fits']
def __init__(self, root, transform=None, target_transform=None,
loader=None):
if loader is None:
loader = self.__fits_loader
super(FitsFolder, self).__init__(root, loader, self.EXTENSIONS,
transform=transform,
target_transform=target_transform)
@staticmethod
def __fits_loader(filename):
file = fits.open(filename)
return (file[1].data)
But when I implement the actual loading process of data:
testset = FitsFolder(root='/Documents/Image_data2')
test_loader = torch.utils.data.DataLoader(testset, batch_size=batch_size,shuffle=False)
And try to plot them, the images look nothing like they are supposed to:
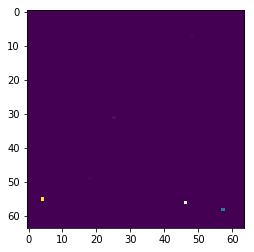

Rather than:
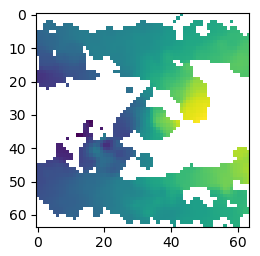
I also don’t know how to perform a transform such as:
transform = transforms.Compose([
transforms.Resize((32,32)),
transforms.ToTensor(),
])
When using this setup…any help would be greatly appreciated!
Could you attach an example file and a converted image (PNG or JPG) for me to verify that my method works?
EDIT: from what I just saw from googling: it should work with
img_data = fits.get_data(filename)
As load function and then passing the numpy array to the transformations
If you take a look at the gist I linked in the other post:
data.Dataset comes from import torch.utils.data as data
Unfortunately if I use .get_data I get the warning: module 'astropy.io.fits' has no attribute 'get_data' so I’m currently having to manually pull out the data using:
def __fits_loader(filename):
file = fits.get_data(filename)
return file
So this method is not quite working yet.
I would upload a FITS example but they are not valid extensions for this forum sorry.
Okay that would be really helpful, thanks! I’ve uploaded them to a git repo here:
and added the FITS file to Archive if that’s easier.
Ah, it works if I use getdata() rather than get_data(), so you were right. Although interestingly I have to multiply the output of using getdata() by 1 to make the image not throw up warnings.
1 Like
Strange. If I try use fits.get_data() in a jupyter notebook it works like a charm but if I want to use it within a function in a python-file it breaks.
However I got the following code working with your fits-file by surpassing the get_data() function:
link_to_gist
The code is usually straight forward. If you have any questions regarding the functions feel free to ask.
The output I got seems pretty similar to your uploaded image:

1 Like
Really strange isn’t it!
Thanks it works great and gives the same image as what I was seeing with my more crude method, thanks!
I’ve noticed in default_fits_loader you have left a space for a custom label function. Would it be possible to alter the loader so that instead of loading single fits files, it would load a stacked fits data cube, split the 2d arrays and assign them the labels?
This would help to save a lot of space by saving cubes instead of lots of fits files in preprocessing.
should be possible, if the labels are saved in the same order as the arrays inside the cube
Hi,
Can you provide a repo link or explain how you managed transform the data using Compose. I am stuck on normalizing and transforming the FITS image
Hi @Shoebhabeeb, sorry for the delayed reply! I’ve just seen this, as it’s been a while since I encountered this problem I’ll need to go back and have a look so I’ll get back to you on this as soon as I can… it could be that using Compose isn’t the optimal way to do this anymore so I’ll check that first 
Hi,
Sure… Do let me know as soon as you can.
I am having trouble passing my dataloader into transforms functions. As they usually classify my images as Nonetype



