Hi all,
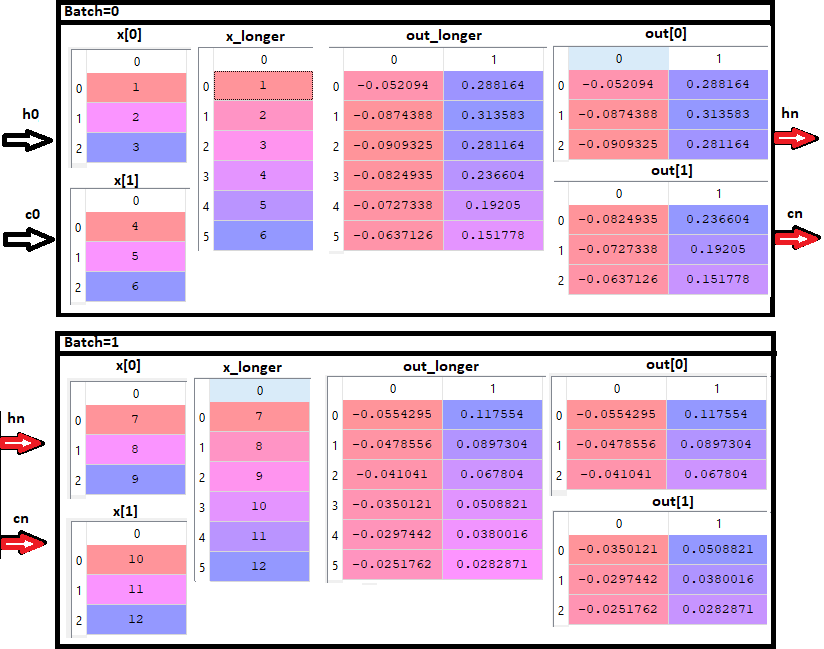
I’m trying to train a network with LSTMs to make predictions on time series data with long sequences. The sequence length is too long to be fed into the network at once and instead of feeding the entire sequence I want to split the sequence into subsequences and propagate the hidden state to capture long term dependencies. I’ve done this successfully before with Keras passing the ‘stateful=True’ flag to the LSTM layers, but I’m confused about how to accomplish the same with PyTorch.
In particular, I’m not sure how to keep and propagate the hidden states when feeding subsequences of a longer sequence as batches. What i’ve tried so far is to do something like:
def forward(self, batch_data):
self.hidden = [Variable(h.data) for h in self.hidden]
lstm_out, self.hidden = self.lstm1(batch_data, self.hidden)
y_pred = self.sigmoid(self.fc1(lstm_out[:,-1]))
to maintain the hidden state values, between batches and then set them to zero when starting on a new sequence.
I’ve written up a notebook to illustrate what I’m to trying achieve: https://github.com/shaurya0/pytorch_stateful_lstm/blob/master/pytorch_stateful_lstm.ipynb
The model is based off of http://philipperemy.github.io/keras-stateful-lstm/