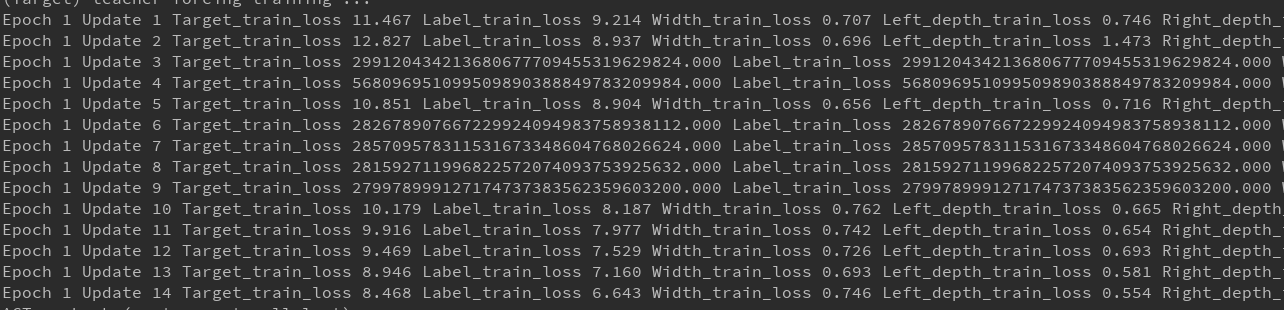
When I use cross entropy loss in my code, with nn.NLLLoss() or code implemented by myself, the loss is very strange, like the picture. Could it be the problem with my pytorch version?
Could you post some information about the last non-linearity your model is using and also with criterion you are applying?
Note that nn.CrossentropyLoss expects raw logits, while nn.NLLLoss needs log probabilities calculated by F.log_softmax.
Also, could you check the input and output of your model for the high loss values?
The last non-linearity is F.tanh()
the details of criterion I use are attached below
def forward(self, input, target, mask):
# truncate to the same size
# input, target, mask like (60, 17, 2631), (60, 17), (60, 17)
target = target[:, :input.size(1)]
mask = mask[:, :input.size(1)]
"""
batch_size = target.size(0)
seq_length = target.size(1)
input = input.view(batch_size * seq_length, -1)
target = target.view(batch_size * seq_length)
# mask = mask.view(batch_size)
output = self.loss(input, target)
output = output.view(batch_size, seq_length)
output = output * mask
"""
# pick the dims correspond to words
output = -input.gather(2, target.unsqueeze(2)).squeeze(2) * mask # like (60, 17)
output = torch.sum(output) / torch.sum(mask) # scalar, average log likelihoods across all time steps of all captions in a batch, positive values
return output
I both tried to use nn.NLLLoss or use gather to calculate loss by myself, but neither of them works. The input of this function is after F.log_softmax().
Actually, I want to supplement is that the same code but with a different dataset will work well. However, I do not think it is dataset’s fault, because the same dataset I use under other code works.
So you are applying F.log_softmax(F.tanh(output), 1) on the model outputs?
How do you calculate the mask?
Since you are dividing by the sum of it, it looks like a good starting point to look for exploding loss values.
Morning,
I am trying to use the cross entropy loss function but i keep getting the error:
RuntimeError: the derivative for 'target' is not implemented
What is causing this? Do i need to perform an operation on the input data?? Can anyone help?
chaslie
Could you post a code snippet showing how you use this method? Usually you would just use criterion(output, target), where target does not require a gradient.
hi ptrblck,
The code is as follows:
def loss_fn(out2, data_loss, z_loc, z_scale):
BCE = F.cross_entropy(data_loss,out2, size_average=True,reduction ='mean')
elsewhere in the code:
data = data.cuda()
z_loc, z_scale = model.Encoder(data)
out = model.Decoder(z)
data_loss=data
data_loss=data_loss.squeeze(1)
out2=out.squeeze(1)
I am using the squeeze functions because i had an error about a size 3 tensor been used from the output of my network, when cross entropy only needs a size 2 tensor. I have tried it with and wthout the squeeze on the input data and it dosen’t complain…
chaslie
Try to swap data_loss for out2, as the method assumes the output of your model as the first argument and the target as the second.
Ptrblck
changing BCE to:
BCE = F.cross_entropy(out2, data_loss,size_average=True,reduction ='mean')
I get a different error:
RuntimeError: Expected object of scalar type Long but got scalar type Float for argument #2 'target'
I have also removed the squeeze command, but this didn’t help. Am i right in assuming that i have to reformat the data_loss term to long as opposed to float, but this doesn’t make sense, does it?
F.cross_entropy expects a target as a LongTensor containing the class indices.
E.g. for a binary classification use case your output should have the shape [batch_size, nb_classes], while the target should have the shape [batch_size] and contain class indices in the range [0, nb_classes-1].
You could alternatively use nn.BCEWithLogitsLoss or F.binary_cross_entropy_with_logits, which expects a target of the same shape as the model output as a FloatTensor.
Brilliant that works.
Thanks for solving another case of user error/ignorance 
Another Silly question,
when i use MSE loss I get errors between 0 and 1, however the exact same network with binary_cross_entropy_with_logits yields errors -3e+21, why is this? I am guessing its the way the loss function is calculated. Should i use a sigmoid function to normalise the results?
Also looking at the results the output is significantly worse than the MSE loss output, so the second part of the question is, how to choose the loss function ?
No, F.binary_cross_entropy_with_logits expects logits, so applying a sigmoid will yield wrong results.
The statistics between BCE and MSE losses are quite different, so you can’t expect loss values in the same range.
Usually you won’t use MSELoss for a classification use case, but a regression one.
A multi-class classification use case will most likely work fine with nn.CrossEntropyLoss etc.
There are some baseline loss function, which should just work for the specific use cases, but of course there are exceptions as always.
That being said, your loss using BCE looks a bit strange. Could you post the output and target which creates this low loss value?
thanks for the response, clears up the some of the confustion, but not all…
I found the problem, this was partly due to the way the stats were being calculated with the loss, having rewritten that section I am now starting to get better (though not entirely correct) results…
I still faced problem with the F.cross_entropy function. My target/label can be 0 or 1. The shapes should be correct, but I don’t know why the target cannot be 1 when I run the F.cross_entropy function. Any hints to me?
input = torch.tensor([[-127050.7500],
[ 214151.2344],
[ 282199.3125],
[ 291863.4375],
[ 256611.0469],
[ 275604.2188],
[ 429206.9688],
[ -35253.5273],
[ 167581.2969],
[ -27991.6875],
[ -49363.4688],
[ -39173.5508],
[ -42769.9766],
[ -71630.0938],
[ 288389.4375],
[ -99099.6641]])
target = torch.tensor([0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1], dtype=torch.int64)
loss = F.cross_entropy(input, target)
loss
IndexError Traceback (most recent call last)
<ipython-input-79-a109ce392e4a> in <module>
19 #input = torch.randn(3, 5, requires_grad=True)
20 target = torch.tensor([0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1], dtype=torch.int64)
---> 21 loss = F.cross_entropy(input, target)
22 loss
/opt/conda/lib/python3.7/site-packages/torch/nn/functional.py in cross_entropy(input, target, weight, size_average, ignore_index, reduce, reduction)
2315 if size_average is not None or reduce is not None:
2316 reduction = _Reduction.legacy_get_string(size_average, reduce)
-> 2317 return nll_loss(log_softmax(input, 1), target, weight, None, ignore_index, None, reduction)
2318
2319
/opt/conda/lib/python3.7/site-packages/torch/nn/functional.py in nll_loss(input, target, weight, size_average, ignore_index, reduce, reduction)
2113 .format(input.size(0), target.size(0)))
2114 if dim == 2:
-> 2115 ret = torch._C._nn.nll_loss(input, target, weight, _Reduction.get_enum(reduction), ignore_index)
2116 elif dim == 4:
2117 ret = torch._C._nn.nll_loss2d(input, target, weight, _Reduction.get_enum(reduction), ignore_index)
IndexError: Target 1 is out of bounds.
Your input shape is [batch_size=16, 1], which is wrong for a 2-class classification.
nn.CrossEntropyLoss expects the model output to have the shape [batch_size, nb_classes] for a multi-class classification, so in your case this would be [batch_size, 2].
If you want to use a single output neuron for the binary classification, you could use nn.BCEWithLogitsLoss instead (and would need to unsqueeze dim1 in your target).
It seems that your suggestion works! Thank you @ptrblck