Hi,
I’m trying to define a custom Dataset for my medical images. For each patient, I have a histopathology Whole Slide Images (WSI). I preprocess each image, obtaining a variable number of 224 x 224 patches and keep them in a folder with the patient ID. The label for each patch in a given patient folder is the label assigned to the WSI. The data directory structure is as follows:
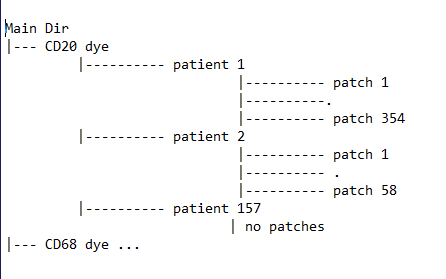
I want to write a custom Dataset that will load each patch, with the correct label and feed it into DataLoader, in this case treating each patch as independent. My current implementation return a list of images and list of labels, as so:
class histoDataset(Dataset):
def __init__(self, main_dir, csv_file, transform): self.main_dir = main_dir self.transform = transform self.CSV = pd.read_csv(csv_file) self.folder_id_code = self.CSV['Patient image ID'].tolist() self.image_diagnosis = self.CSV['Classification.based.on.CD20'].tolist() self.classes = list(dict.fromkeys(self.image_diagnosis)) self.label = self.CSV['Classification.based.on.CD20'].apply(self.classes.index) self.label_dict = dict(zip(self.label, self.classes)) self.folders = [f.path for f in os.scandir(self.main_dir)] def __len__(self): return len(self.folders) def __getitem__(self, idx): image_list = [] image_labels = [] self.folder_loc = os.path.join(self.main_dir, self.folder_id_code[idx]) self.folder_files = [f.path for f in os.scandir(self.folder_loc)] for image in self.folder_files: image = Image.open(image) image_tensor = self.transform(image) image_label = self.label[idx] image_list.append(image_tensor) image_labels.append(image_label) return image_list, image_labels
After inputting the resulting histoDataset into DataLoder and running this:
sample_batch = next(iter(df_loader))
sample_images, sample_labels = sample_batch
print_images = sample_images[0:10]
print_labels = sample_labels[0:10]
to check the images and labels, I get the following error:
Traceback (most recent call last):
File "C:\Users\AmayaGS\Documents\PhD\Data\Dataset_loader.py", line 248, in <module>
sample_batch = next(iter(df_loader))
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\dataloader.py", line 521, in __next__
data = self._next_data()
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\dataloader.py", line 561, in _next_data
data = self._dataset_fetcher.fetch(index) # may raise StopIteration
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\_utils\fetch.py", line 47, in fetch
return self.collate_fn(data)
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\_utils\collate.py", line 84, in default_collate
return [default_collate(samples) for samples in transposed]
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\_utils\collate.py", line 84, in <listcomp>
return [default_collate(samples) for samples in transposed]
File "C:\ProgramData\Anaconda3\lib\site-packages\torch\utils\data\_utils\collate.py", line 82, in default_collate
raise RuntimeError('each element in list of batch should be of equal size')
RuntimeError: each element in list of batch should be of equal size
Could someone please point out how to correctly implement the histoDataset and why I am getting this error? Thanks!