Thank you so much.
I managed to make it work  Here is what I have:
Here is what I have:
class dynamic_dataset(Dataset):
def __init__(self, WS):
self.len = 500
self.WS = WS
def __getitem__(self, index):
image = torch.randn(self.WS,38)
label = torch.randint(1, (1,))
return image, label
def __len__(self):
return self.len
class dynamic_model(nn.Module):
def __init__(self, H_in, W_in, num_kernels):
super(dynamic_model, self).__init__()
C_in_1, C_out_1 = 1, num_kernels
kernel_size_1 = 3
H_out_1, W_out_1 = self.conv_output_shape((H_in, W_in), kernel_size=kernel_size_1) # W_in = 38
C_in_2, C_out_2 = C_out_1, num_kernels
kernel_size_2 = 1
H_out_2, W_out_2 = self.conv_output_shape((H_out_1, W_out_1), kernel_size=kernel_size_2)
self.layer1 = nn.Sequential(
nn.Conv2d(C_in_1, C_out_1, kernel_size=kernel_size_1, stride=1, padding=0),
nn.ReLU())
self.layer2 = nn.Sequential(
nn.Conv2d(C_in_2, C_out_2, kernel_size=kernel_size_2, stride=1, padding=0),
nn.ReLU())
self.fc1 = nn.Linear(C_out_2 * H_out_2 * W_out_2, C_out_2 * H_out_2 * W_out_2)
self.fc2 = nn.Linear(C_out_2 * H_out_2 * W_out_2, 2)
def forward(self, x):
x = x.unsqueeze(1)
x = self.layer1(x)
x = self.layer2(x)
x = x.view(x.size(0), -1)
x = self.fc1(x)
x = self.fc2(x)
return x
def conv_output_shape(self, h_w, kernel_size=1, stride=1, pad=0, dilation=1):
from math import floor
if type(kernel_size) is not tuple:
kernel_size = (kernel_size, kernel_size)
h = floor( ((h_w[0] + (2 * pad) - ( dilation * (kernel_size[0] - 1) ) - 1 )/ stride) + 1)
w = floor( ((h_w[1] + (2 * pad) - ( dilation * (kernel_size[1] - 1) ) - 1 )/ stride) + 1)
return h, w
# simplified experiment setup
if __name__ == '__main__':
lr = 0.001
WS = 25
num_epochs = 3
batch_size = 128
num_kernels = 32
model = dynamic_model(WS, 38, num_kernels)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=lr)
trainset = dynamic_dataset(WS=WS)
trainloader = DataLoader(trainset, batch_size=batch_size)
for i, (images, labels) in enumerate(trainloader):
labels = labels.squeeze(1)
optimizer.zero_grad()
outputs = model(images)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
print(i, loss.item())
Sorry for the long code snippet. Could you please point out anything in my model structure that I could improve on? Layer 1 and layer 2 kernel_size?
Really appreciate all your help.
Thank you 
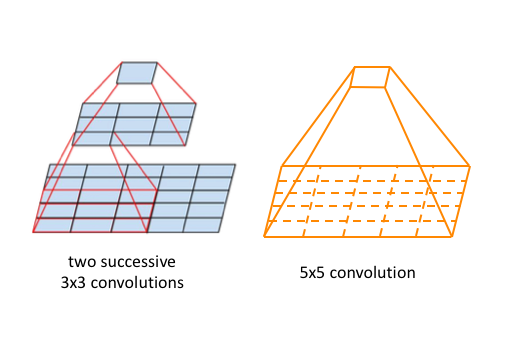
 . A traditional CNN has fixed kernel sizes, so that you can train every weight at the same time. This ensures that the model is consistent. Assuming that you have a maximum number of kernels and size of kernels, then if you train only part of the kernels and parts of the kernels, it probably breaks the global behaviour of the model.
. A traditional CNN has fixed kernel sizes, so that you can train every weight at the same time. This ensures that the model is consistent. Assuming that you have a maximum number of kernels and size of kernels, then if you train only part of the kernels and parts of the kernels, it probably breaks the global behaviour of the model.
 Here is what I have:
Here is what I have: