Hello,
I am using the following codes to finding the class probabilities to classify channels into different labels. The training was of 6 class labels[0,1,2,3,4,56] in a one-hot encoded method.
for bIdx, sample in enumerate(test_loader):
img, seg = sample[0].to(device), sample[1].to(device) # shape --> [2,6,48,64,64]
pred = model(img)
lbl_pred = pred.data.max(1)[1].cpu().numpy()[:, :, :].astype('uint8')
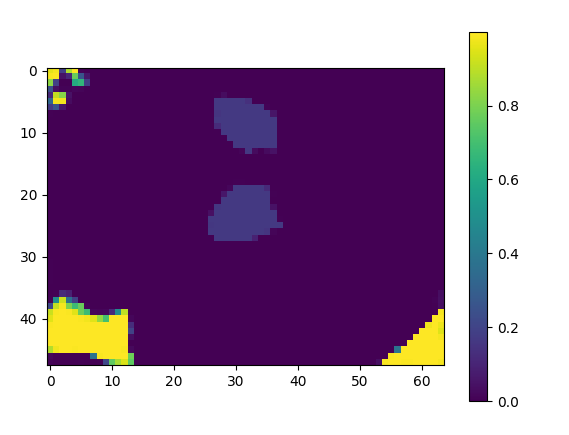
But I am missing label 5 in the prediction/output(the cursor in the following image).

This shows as label 0 which is clearly not the case as label 0: background.
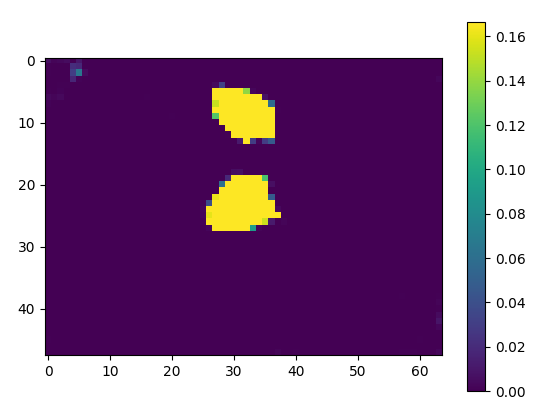
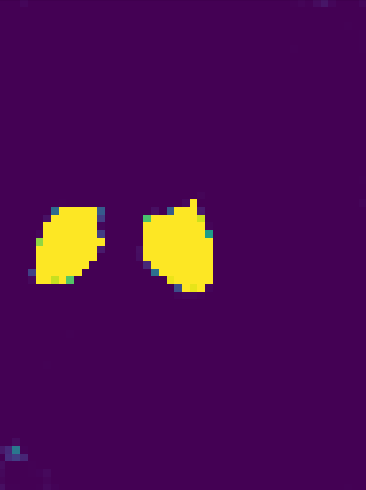
But if I check pred.data separately, it has label 5 probability values in it.
plt.imshow(pred.data[0, 5, :, 20, :].cpu().numpy())
plt.show()
resulting in the following image.

Am I missing anything?