Hi, if I have come to the wrong discussion board, please forgive me and let me know the proper one.
I need your feedback on my model and training process.
I was trying to learn the GAT model using multiple graph datasets at once.
However, there were data imbalances between positive and negative labels across the dataset.
for i in range(len(GraphList)):
print(torch.bincount(GraphList[i].edge_label))
tensor([3292, 1613]) #[the number of 0 labels, the number of 1 labels]
tensor([1724, 1177])
tensor([1532, 1560])
tensor([13707, 5585])
tensor([188, 274])
tensor([2077, 465])
tensor([2092, 608])
tensor([2170, 536])
tensor([2398, 463])
For your understanding, I attach the example of mini-batch data after calling LinkNeighborLoader which is the edge-sampling version of DataLoader.
Data(x=[2], edge_index=[2, 453], edge_label=[32], edge_label_index=[2, 32], edge_class=[32], num_nodes=513, n_id=[513], e_id=[453], batch=[513], num_sampled_nodes=[3], num_sampled_edges=[2], input_id=[32], edge_index_class=[453])
To train the model classifying the binary labels for all datasets, I constructed the model as below which is based on the pytorch geometric.
class MyModel(torch.nn.Module):
def __init__(self, in_channels, hidden_channels, out_channels):
super(MyModel, self).__init__()
# MLP
self.convert_layer = nn.Sequential(
nn.Linear(4, 16, bias=True),
nn.LeakyReLU(),
nn.Linear(16, 64, bias=True),
nn.LeakyReLU(),
nn.Linear(64, 256, bias=True)
)
# GAT
self.conv1 = GATConv(in_channels, hidden_channels, heads=6) # in_channel, hidden_channel, heads
self.lin1 = torch.nn.Linear(in_channels, hidden_channels * 6)
self.norm1 = LayerNorm(hidden_channels * 6)
self.conv2 = GATConv(hidden_channels * 6, out_channels, heads=1, concat=False) # hidden_channel * heads, out_channel, heads
self.lin2 = torch.nn.Linear(hidden_channels * 6, out_channels)
def convert_data_size(self, data1, data2):
x2 = torch.zeros((len(data1), 256), dtype=torch.float).to(device) #(107940, 256)
x3 = self.convert_layer (data2).to(device) #(19392, 256)
nonzero_index = torch.tensor(cID).to(device)
x2.index_add_(0, nonzero_index, x3) #(107940, 256)
return x2
def forward(self, data1, data2, neighbor_cl_ids, edge_index):
x_dict, attn1_dict, attn2_dict = {}, {}, {}
for i in range(len(data2)):
data2[i] = self.convert_data_size(data1, data2[i]) #(107940, 256), requires_grad=true
for e_i_class in torch.unique(neighbor_cl_ids):
x = torch.cat((data1, data2[int(e_i_class.item())]), dim=1) #(107940, 512)
x_1, a1 = self.conv1(x, edge_index, return_attention_weights=True)
x = F.leaky_relu(self.norm1(x_1))
x = F.dropout(x, p = 0.2, training = self.training)
x_2, a2 = self.conv2(x, edge_index, return_attention_weights=True)
x = x_2 + self.lin2(x)
x_dict[int(e_i_class.item())] = x
attn1_dict[int(e_i_class.item())] = a1
attn2_dict[int(e_i_class.item())] = a2
return x_dict, attn1_dict, attn2_dict
for epoch in range(1, num_epochs+1):
tr_losses = 0
val_losses = 0
tr_loss_sum = 0
val_loss_sum = 0
model.train()
for data in tqdm(train_loader):
tr_loss = []
data = data.to(device)
data.edge_class = data.edge_class[data.input_id]
data.edge_index_class = torch.zeros(len(data.edge_index[0])).to(device)
for i in range(len(data.edge_index_class)):
if ((data.edge_index[1][i] in data.edge_label_index[0]) or (data.edge_index[1][i] in data.edge_label_index[1])):
data.edge_index_class[i] = data.edge_class[(data.edge_index[1][i]==data.edge_label_index[0])
|(data.edge_index[1][i]==data.edge_label_index[1])]
else:
data.edge_index_class[i] = torch.max(data.edge_index_class[(data.edge_index[1][i]==data.edge_index[0])])
optimizer.zero_grad()
z, a1, a2 = model(data.x[0], data.x[1], data.edge_index_class, data.edge_index)
for i in torch.unique(data.edge_class):
tr_loss_mean = 0
i = int(i.item())
try:
tr_out = ((z[i][data.edge_label_index[0][data.edge_class==i]] * z[i][data.edge_label_index[1][data.edge_class==i]]).mean(dim=-1)).view(-1)
tr_loss.append(criterion(tr_out, data.edge_label[data.edge_class==i].float()))
except KeyError:
pass
tr_loss_mean = sum(tr_loss)/len(tr_loss)
tr_loss_mean.backward()
optimizer.step()
tr_losses += tr_loss_mean.item()
avg_tr_loss = tr_losses/len(train_loader.dataset)
model.eval()
with torch.no_grad():
y_val_pred, y_val_pred_prob, y_val_true = [], [], []
for data in tqdm(val_loader):
val_loss = []
data = data.to(device)
data.edge_class = data.edge_class[data.input_id]
data.edge_index_class = torch.zeros(len(data.edge_index[0])).to(device)
for i in range(len(data.edge_index_class)):
if ((data.edge_index[1][i] in data.edge_label_index[0]) or (data.edge_index[1][i] in data.edge_label_index[1])):
data.edge_index_class[i] = data.edge_class[(data.edge_index[1][i]==data.edge_label_index[0])
|(data.edge_index[1][i]==data.edge_label_index[1])]
else:
data.edge_index_class[i] = torch.max(data.edge_index_class[(data.edge_index[1][i]==data.edge_index[0])])
y_val_true.append(data.edge_label)
z, a1, a2 = model(data.x[0], data.x[1], data.edge_index_class, data.edge_index)
for i in torch.unique(data.edge_class):
val_loss_mean = 0
i = int(i.item())
try:
val_out = ((z[i][data.edge_label_index[0][data.edge_class==i]] * z[i][data.edge_label_index[1][data.edge_class==i]]).mean(dim=-1)).view(-1)
val_out_sig = ((z[i][data.edge_label_index[0][data.edge_class==i]] * z[i][data.edge_label_index[1][data.edge_class==i]]).mean(dim=-1)).view(-1).sig
val_loss.append(criterion(val_out, data.edge_label[data.edge_class==i].float()))
except KeyError:
pass
y_val_pred.append((val_out_sig>0.5).float().cpu())
y_val_pred_prob.append((val_out_sig).float().cpu())
val_loss_mean = sum(val_loss) / len(val_loss)
val_losses += val_loss_mean.item()
avg_val_loss = val_losses/len(val_loader.dataset)
print(f'Epoch: {epoch:03d}, Training Loss: {avg_tr_loss:.4f}, Validation Loss: {avg_val_loss:.4f}')
y, pred, pred_prob = torch.cat(y_val_true, dim=0).cpu().numpy(), torch.cat(y_val_pred, dim=0).cpu().numpy(), torch.cat(y_val_pred_prob, dim=0).cpu().numpy()
for i in torch.unique(val_loader.data.edge_class):
i = int(i.item())
val_f1 = f1_score(y[val_loader.data.edge_class==i], pred[val_loader.data.edge_class==i]) #average='micro'
val_auc = roc_auc_score(y[val_loader.data.edge_class==i], pred_prob[val_loader.data.edge_class==i])
val_aupr = average_precision_score(y[val_loader.data.edge_class==i], pred_prob[val_loader.data.edge_class==i])
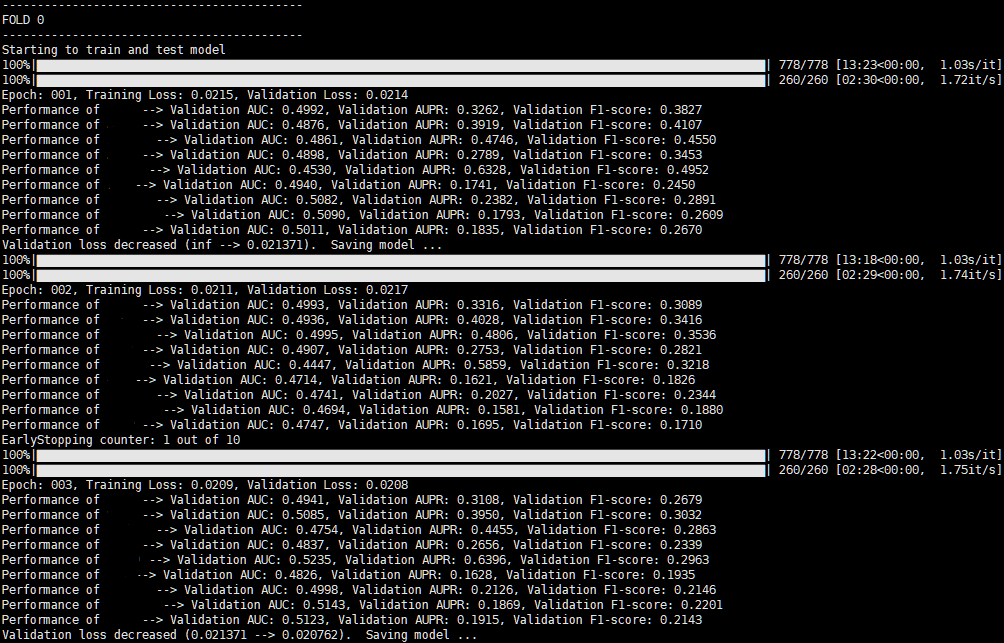
print(f'Performance of {cellline_list[i]} --> Validation AUC: {val_auc:.4f}, Validation AUPR: {val_aupr:.4f}, Validation F1-score: {val_f1:.4f}')
After that, I got the result with very poor AUC, AUPR, and F1 scores as below.
Unfortunately, the scores continued to drop as each epoch passed.
I expected that the model could learn the pattern of positive and negative samples by sharing the same parameters in conv1 and conv2, but it looks failed.
If there are not only data imbalance problems but also training processes, please tell me.
Thank you for reading the question.
Have a nice day!
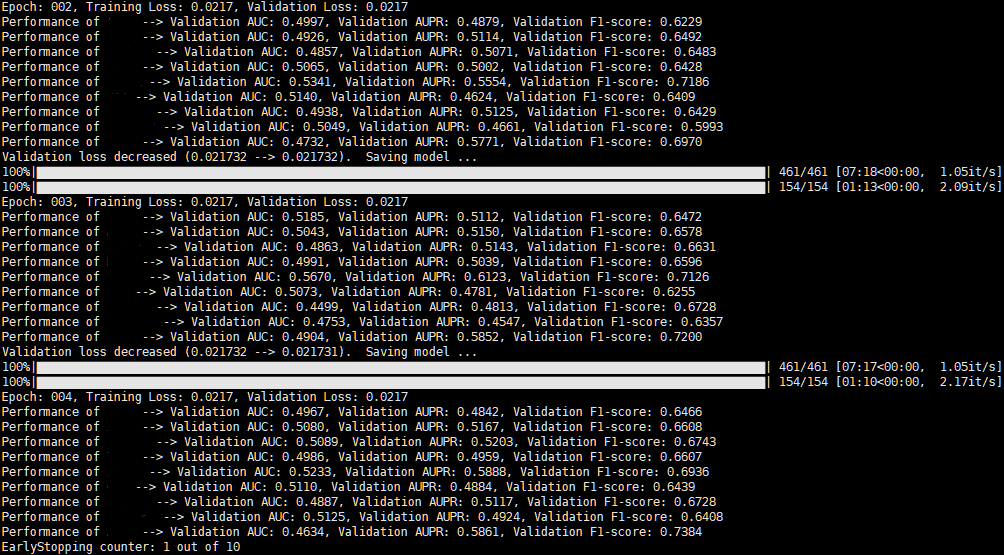
[Update]
I tried to balance the ratio of the positive and negative samples, but it still had poor performance.
Are there any possible approaches to make the model learn?