Hello All,
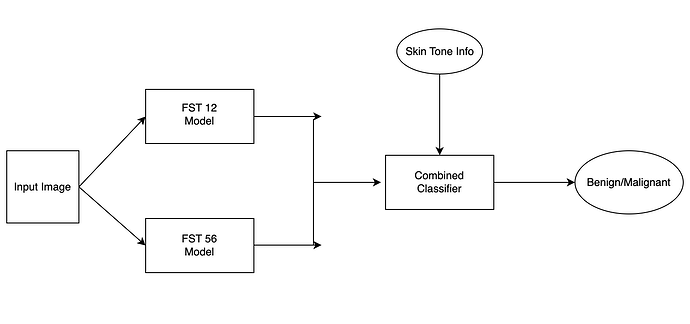
I am working with the Stanford DDI Dataset. Basically, the data contains images of dermatology related diseases classified as either Benign or Malignant. Furthermore, the data is diversified across various skin tones (FST 12 - fair skin, FST 56- dark skin, FST 34 -In Between). Without the skin tone information, the problem is fairly binary classification (Either benign or malignant). However, I want to build a model which consists of two pre-trained models (One trained on FST 12, One trained on FST 56) based on this idea and then have them output the label benign/malignant. I am not sure how to structure the data in folders though. This is what I want to build.
Any help is appreciated.
Thanks!
Please take a look at this: https://pytorch.org/vision/stable/generated/torchvision.datasets.ImageFolder.html
If you end up using ImageFolder then you can create an ensemble of models (similar to your proposed solution) where each model will train/predict using different dataset located in different folders i.e.
FST_12/train/benign_class_images (for training dataset)
FST_12/train/malignant_class_images (for training dataset)
FST_12/test_images
FST_56/train/benign_class_images (for training dataset)
FST_56/train/malignant_class_images (for training dataset)
FST_56/test_images
Depending on how the images are currently named, you can automate the move operations using a python/shell script.
Hey,
Thanks for the suggestion. I have used the ImageFolder approach and trained the models separately (also while following a similar folder structure as you specified).
I have also saved the state_dicts for these models and I am able to load them in to the ensemble model. Now, I was wondering how I could inject the skin_type info to the outputs of both these models and pass them through some layers before making a final classification?
Thank you
@Manny_Parkinson, Sounds like you want to separate the images based on skin_type before predicting benign/malignant. Maybe you can train a model to perform binary classification on skin_type then separate the images into the folder structures mentioned previously and finally use the existing ensemble (the one you already trained) to predict benign/malignant.
@tiramisuNcustard, I don’t necessarily want to separate the images based on skin type. I just want to provide an auxiliary input, (the skin tone) to have some sort of weightage when making a vote. I apologize if I cannot articulate it better. For now, I was able to train two models individually based on this directory structure.
12/train/Benign
12/train/Malignant
12/val/Benign
12/val/Malignant
The same for 56 skin tone as well.
What I am struggling at is training both the models together while also providing an additional input about the skin tone. The problem arises when thinking about how to structure the data for combined training and how to provide the auxiliary input label (skin tone) as the model trains.
I really really appreciate your patience.
@Manny_Parkinson, Sounds like you can differentiate between the skin tone based on the images alone, so I am assuming the images are color images. Models are capable of extracting features from all 3 channels (red, green and blue) of color images. Skin tones will lead to different signatures (i.e. characteristic variation in one or more of these channels) in these images. This means that the information about the skin tone is already present in all the images. I would only create a binary classifier (i.e. benign or malignant) and disregard the skin tone classification as the presence/absence of the disease is the constant to be determined across all skin tones.
If you must, add text annotations on all the images manually before training. Regards.