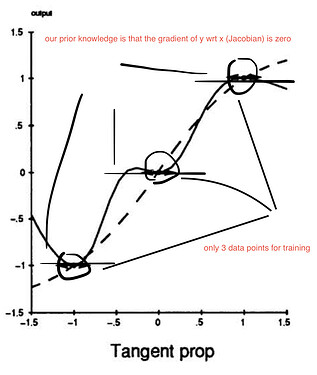
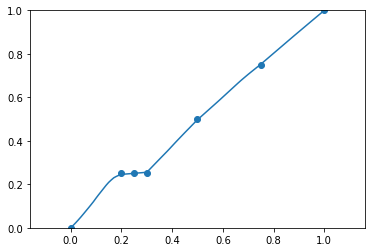
I’m trying to implement an old paper: Tangent Prop - A formalism for specifying selected invariances in an adaptive network. .
For example, let’s consider the following regression problem. Suppose we have 1-dimensional input (x) and 1-dimensional output (y). In this case, the Jacobian matrix is just 1-by-1 (a scalar) that describes the derivative of y w.r.t. x. So far so good.
So we iteratively perform gradient descent for the regression sum-of-squares loss + adjustable parameter * the squared Jacobian. Ideally, when this loss is minimized, we get the learned function as the solid line (see diagram below) given that our neural net has enough capacity.
Without the squared Jacobian term, we would get the learned function as the dashed line (see diagram below).
However, what I described above fails when implemented in PyTorch as follows:
class Net(nn.Module):
def __init__(self):
super().__init__()
self.main = nn.Sequential(
nn.Linear(1, 100),
nn.ReLU(),
nn.Linear(100, 100),
nn.ReLU(),
nn.Linear(100, 1)
)
def forward(self, x):
return self.main(x)
def get_model():
model = Net()
return model, optim.Adam(model.parameters(), lr=1e-3)
def sos(ypred, y):
"Sum of squares loss"
return torch.sum((ypred - y) ** 2)
def get_jacobian(net, x, noutputs):
x.requires_grad = True
y = net(x)
grad_params = torch.autograd.grad(y, x, create_graph=True)
return grad_params[0]
# just a straight line with slope 1 and intercept 0
data = [0., 0.25, 0.5, 0.75, 1.0]
labels = [0., 0.25, 0.5, 0.75, 1.0]
model, opt = get_model()
jacobian_losses = []
reg_losses = []
for i in range(100):
ypred = model(torch.tensor(data).view(-1, 1))
reg_loss = sos(ypred, torch.tensor(labels).view(-1, 1))
reg_losses.append(float(loss))
jacobian_loss = 0
for x in [0., 0.25, 0.5, 0.75, 1.0]:
jacobian = get_jacobian(model, torch.tensor([[x]]), 1)
temp = torch.norm(jacobian) ** 2
jacobian_loss += temp
jacobian_losses.append(float(jacobian_loss))
total_loss = reg_loss + 1e-1 * jacobian_loss
total_loss.backward()
opt.step()
opt.zero_grad()
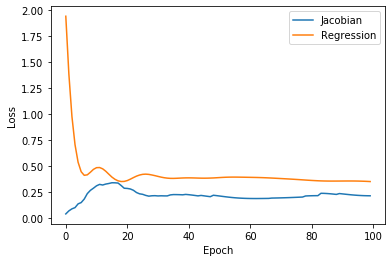
Here’s the losses plotted together:
plt.plot(jacobian_losses, label='Jacobian')
plt.plot(reg_losses, label='Regression')
plt.xlabel('Epoch'); plt.ylabel('Loss')
plt.legend()
plt.show()
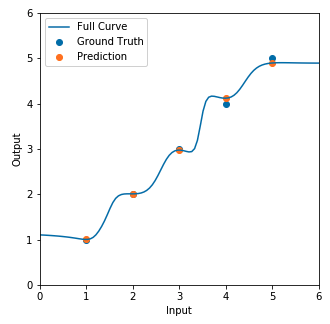
Here’s the resulting regression line:
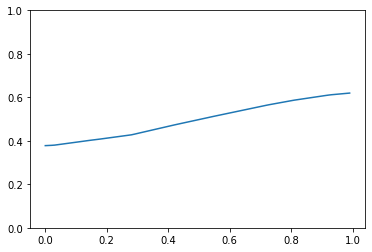
And we don’t see the polynomial-like regression line like we saw above in the figure given by the paper. Can anyone help point out my mistake(s)? I’ve tried multiplying Jacobian loss by a smaller weight but I still don’t see the curvatures.