I am wondering if pytorch has gaussian filtering (convolution).
For example, if I want to do low pass Gaussian filter on an image, is it possible?
In other words, Im trying to do this:
scipy.ndimage.gaussian_filter(a,b)
https://docs.scipy.org/doc/scipy/reference/generated/scipy.ndimage.gaussian_filter.html
You can create a Conv2d layer and specify the weights to be gaussian.
Then just apply the conv layer on your image.
For anyone who has a problem implementing this here is a solution entirely written in pytorch:
# Set these to whatever you want for your gaussian filter
kernel_size = 15
sigma = 3
# Create a x, y coordinate grid of shape (kernel_size, kernel_size, 2)
x_cord = torch.arange(kernel_size)
x_grid = x_cord.repeat(kernel_size).view(kernel_size, kernel_size)
y_grid = x_grid.t()
xy_grid = torch.stack([x_grid, y_grid], dim=-1)
mean = (kernel_size - 1)/2.
variance = sigma**2.
# Calculate the 2-dimensional gaussian kernel which is
# the product of two gaussian distributions for two different
# variables (in this case called x and y)
gaussian_kernel = (1./(2.*math.pi*variance)) *\
torch.exp(
-torch.sum((xy_grid - mean)**2., dim=-1) /\
(2*variance)
)
# Make sure sum of values in gaussian kernel equals 1.
gaussian_kernel = gaussian_kernel / torch.sum(gaussian_kernel)
# Reshape to 2d depthwise convolutional weight
gaussian_kernel = gaussian_kernel.view(1, 1, kernel_size, kernel_size)
gaussian_kernel = gaussian_kernel.repeat(channels, 1, 1, 1)
gaussian_filter = nn.Conv2d(in_channels=channels, out_channels=channels,
kernel_size=kernel_size, groups=channels, bias=False)
gaussian_filter.weight.data = gaussian_kernel
gaussian_filter.weight.requires_grad = False
You probably want to keep the size of your input the same after it has been filtered. In that case just add some padding to the input (I use reflection padding).
@tetratrio I assume you have a class when you define it, right? otherwise the self.* does not make sense
Oh yea, I forgot to remove that. Thought it would make the code more readable without the class
Yeah!
Good stuff!
Thanks
This doesn’t work with the latest version of PyTorch (1.0), as the LongTensor objects round values when multiplying by floats and end up with a 0 sum, which blows up the torch.exp (causing Python kernel to restart in Jupyter).
The fix however is easy … ensure xy_grid is a FloatTensor, by changing adding .float() to that one line like this:
#...
xy_grid = torch.stack([x_grid, y_grid], dim=-1).float()
#...
Here is the full function version for PyTorch 1.0:-
# From https://discuss.pytorch.org/t/is-there-anyway-to-do-gaussian-filtering-for-an-image-2d-3d-in-pytorch/12351/3
def get_gaussian_kernel(kernel_size=3, sigma=2, channels=3):
# Create a x, y coordinate grid of shape (kernel_size, kernel_size, 2)
x_coord = torch.arange(kernel_size)
x_grid = x_coord.repeat(kernel_size).view(kernel_size, kernel_size)
y_grid = x_grid.t()
xy_grid = torch.stack([x_grid, y_grid], dim=-1).float()
mean = (kernel_size - 1)/2.
variance = sigma**2.
# Calculate the 2-dimensional gaussian kernel which is
# the product of two gaussian distributions for two different
# variables (in this case called x and y)
gaussian_kernel = (1./(2.*math.pi*variance)) *\
torch.exp(
-torch.sum((xy_grid - mean)**2., dim=-1) /\
(2*variance)
)
# Make sure sum of values in gaussian kernel equals 1.
gaussian_kernel = gaussian_kernel / torch.sum(gaussian_kernel)
# Reshape to 2d depthwise convolutional weight
gaussian_kernel = gaussian_kernel.view(1, 1, kernel_size, kernel_size)
gaussian_kernel = gaussian_kernel.repeat(channels, 1, 1, 1)
gaussian_filter = nn.Conv2d(in_channels=channels, out_channels=channels,
kernel_size=kernel_size, groups=channels, bias=False)
gaussian_filter.weight.data = gaussian_kernel
gaussian_filter.weight.requires_grad = False
return gaussian_filter
Cheers!
Since my last post there has also been the addition of torch.meshgrid. This is what I currently use (it does not contain parameters and works for 1d, 2d and 3d data):
import math
import numbers
import torch
from torch import nn
from torch.nn import functional as F
class GaussianSmoothing(nn.Module):
"""
Apply gaussian smoothing on a
1d, 2d or 3d tensor. Filtering is performed seperately for each channel
in the input using a depthwise convolution.
Arguments:
channels (int, sequence): Number of channels of the input tensors. Output will
have this number of channels as well.
kernel_size (int, sequence): Size of the gaussian kernel.
sigma (float, sequence): Standard deviation of the gaussian kernel.
dim (int, optional): The number of dimensions of the data.
Default value is 2 (spatial).
"""
def __init__(self, channels, kernel_size, sigma, dim=2):
super(GaussianSmoothing, self).__init__()
if isinstance(kernel_size, numbers.Number):
kernel_size = [kernel_size] * dim
if isinstance(sigma, numbers.Number):
sigma = [sigma] * dim
# The gaussian kernel is the product of the
# gaussian function of each dimension.
kernel = 1
meshgrids = torch.meshgrid(
[
torch.arange(size, dtype=torch.float32)
for size in kernel_size
]
)
for size, std, mgrid in zip(kernel_size, sigma, meshgrids):
mean = (size - 1) / 2
kernel *= 1 / (std * math.sqrt(2 * math.pi)) * \
torch.exp(-((mgrid - mean) / (2 * std)) ** 2)
# Make sure sum of values in gaussian kernel equals 1.
kernel = kernel / torch.sum(kernel)
# Reshape to depthwise convolutional weight
kernel = kernel.view(1, 1, *kernel.size())
kernel = kernel.repeat(channels, *[1] * (kernel.dim() - 1))
self.register_buffer('weight', kernel)
self.groups = channels
if dim == 1:
self.conv = F.conv1d
elif dim == 2:
self.conv = F.conv2d
elif dim == 3:
self.conv = F.conv3d
else:
raise RuntimeError(
'Only 1, 2 and 3 dimensions are supported. Received {}.'.format(dim)
)
def forward(self, input):
"""
Apply gaussian filter to input.
Arguments:
input (torch.Tensor): Input to apply gaussian filter on.
Returns:
filtered (torch.Tensor): Filtered output.
"""
return self.conv(input, weight=self.weight, groups=self.groups)
Again, I suggest adding reflection padding.
smoothing = GaussianSmoothing(3, 5, 1)
input = torch.rand(1, 3, 100, 100)
input = F.pad(input, (2, 2, 2, 2), mode='reflect')
output = smoothing(input)
Thank you for the helpful implementation, but I think line:
kernel *= 1 / (std * math.sqrt(2 * math.pi)) * \
torch.exp(-((mgrid - mean) / (2 * std)) ** 2)
should change to
kernel *= 1 / (std * math.sqrt(2 * math.pi)) * \
torch.exp((-((mgrid - mean) / std) ** 2) / 2)
since:
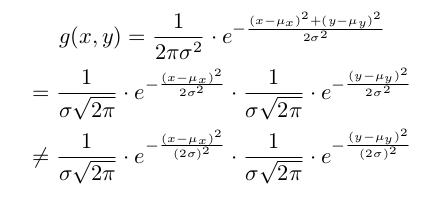
Hey, nice spot there, you’re totally right. I can’t edit my previous post, hope people dont just copy the code without looking at your answer first. Ill post an altered copy here as well
import math
import numbers
import torch
from torch import nn
from torch.nn import functional as F
class GaussianSmoothing(nn.Module):
"""
Apply gaussian smoothing on a
1d, 2d or 3d tensor. Filtering is performed seperately for each channel
in the input using a depthwise convolution.
Arguments:
channels (int, sequence): Number of channels of the input tensors. Output will
have this number of channels as well.
kernel_size (int, sequence): Size of the gaussian kernel.
sigma (float, sequence): Standard deviation of the gaussian kernel.
dim (int, optional): The number of dimensions of the data.
Default value is 2 (spatial).
"""
def __init__(self, channels, kernel_size, sigma, dim=2):
super(GaussianSmoothing, self).__init__()
if isinstance(kernel_size, numbers.Number):
kernel_size = [kernel_size] * dim
if isinstance(sigma, numbers.Number):
sigma = [sigma] * dim
# The gaussian kernel is the product of the
# gaussian function of each dimension.
kernel = 1
meshgrids = torch.meshgrid(
[
torch.arange(size, dtype=torch.float32)
for size in kernel_size
]
)
for size, std, mgrid in zip(kernel_size, sigma, meshgrids):
mean = (size - 1) / 2
kernel *= 1 / (std * math.sqrt(2 * math.pi)) * \
torch.exp(-((mgrid - mean) / std) ** 2 / 2)
# Make sure sum of values in gaussian kernel equals 1.
kernel = kernel / torch.sum(kernel)
# Reshape to depthwise convolutional weight
kernel = kernel.view(1, 1, *kernel.size())
kernel = kernel.repeat(channels, *[1] * (kernel.dim() - 1))
self.register_buffer('weight', kernel)
self.groups = channels
if dim == 1:
self.conv = F.conv1d
elif dim == 2:
self.conv = F.conv2d
elif dim == 3:
self.conv = F.conv3d
else:
raise RuntimeError(
'Only 1, 2 and 3 dimensions are supported. Received {}.'.format(dim)
)
def forward(self, input):
"""
Apply gaussian filter to input.
Arguments:
input (torch.Tensor): Input to apply gaussian filter on.
Returns:
filtered (torch.Tensor): Filtered output.
"""
return self.conv(input, weight=self.weight, groups=self.groups)
smoothing = GaussianSmoothing(3, 5, 1)
input = torch.rand(1, 3, 100, 100)
input = F.pad(input, (2, 2, 2, 2), mode='reflect')
output = smoothing(input)
Another solution would be to directly use PIL’s ImageFilter.GaussianBlur transform.
You can then create the transform:
import numbers
import numpy as np
from PIL import ImageFilter
class GaussianSmoothing(object):
def __init__(self, radius):
if isinstance(radius, numbers.Number):
self.min_radius = radius
self.max_radius = radius
elif isinstance(radius, list):
if len(radius) != 2:
raise Exception(
"`radius` should be a number or a list of two numbers")
if radius[1] < radius[0]:
raise Exception(
"radius[0] should be <= radius[1]")
self.min_radius = radius[0]
self.max_radius = radius[1]
else:
raise Exception(
"`radius` should be a number or a list of two numbers")
def __call__(self, image):
radius = np.random.uniform(self.min_radius, self.max_radius)
return image.filter(ImageFilter.GaussianBlur(radius))
Then you can simply add it to your composed transform before ToTensor():
from PIL import Image
import torchvision.transforms as T
# just add it before ToTensor(), which is ommited here
transform = T.Compose([
GaussianSmoothing([0, 5])
])
TEST_IMG = "path/to/image"
img = Image.open(TEST_IMG)
img_t = transform(img)
img_t.show()
thanks a lot for the code, if you are working with that class through the training - do you need to backprop through it? I mean, I don’t want to change the coefficients just to correctly update other layers before. thanks in advance
No problem, glad you found it useful!
The actual weights are registered as a buffer
self.register_buffer('weight', kernel)
Since they are not parameters (torch.nn.Parameter) they are not trained and will remain static when you use the layer.
class MyModel(torch.nn.Module):
def __init__(self):
super(MyModel, self).__init__()
self.layer = torch.nn.Conv2d(5, 5, kernel_size=3, padding=1)
self.smoothing = GaussianSmoothing(5, kernel_size=3, sigma=1)
def forward(self, x):
x = self.layer(x)
x = torch.nn.functional.pad(x, (1, 1, 1, 1), mode='reflect')
x = self.smoothing(x)
return x
my_model = MyModel()
opt = torch.nn.optim.SGD(my_model.parameters(), lr=1e-3)
output = my_model(torch.randn(1, 5, 10, 10))
loss = output.mean()
opt.zero_grad()
loss.backward()
opt.step()
The smoothing layer will remain static during training
ok. thanks a lot!
just a general thought: If the smoothing operator is the one of the layers in the network - in order to perform backprop - do you need to calculate the derivatives of that operator
you can also do that using kornia.gaussian_blur2d
Hi
Can you explain the working of the code in brief?
Is there any way to adjust the level of denoising in the code?
Hi Salome,
the level of blurring is given by the radius. The higher it is, the blurrier the result is. For a radius of 0, there is no blurring applied. Most of the code deals with the input, which can be either a number (the radius) or a list of two elements, in which case the radius is chosen randomly from this interval.
Okay.  Thanks a lot
Thanks a lot 
@tetratrio
Thanks for the post - I used this and it really helped me.
Just 1 comment:
for size, std, mgrid in zip(kernel_size, sigma, meshgrids):
mean = (size - 1) / 2
kernel *= 1 / (std * math.sqrt(2 * math.pi)) * \
torch.exp(-((mgrid - mean) / std) ** 2 / 2)
# Make sure sum of values in gaussian kernel equals 1.
kernel = kernel / torch.sum(kernel)
Since you do: kernel = kernel / torch.sum(kernel) then there is no reason to divide by:
std * math.sqrt(2 * math.pi)
The moment you normalize the sum to be 1 divisions by a constant (depending or not on the std) will not effect the final result.
Great work - your code taught me alot about how to use conv (1, 2, 3) in pytorch. It was really cryptic before that.
You are correct! It is an unnecessary extra calculation step but it also may reduce confusion about calculating the kernel as it now is a pure normal density function.
But I have to admit I did not think about that when writing the code so its a nice catch!
Also glad you found it useful. I like the idea of implementing convolutional layers and models to be generic in regards to the number of dimensions in the data!