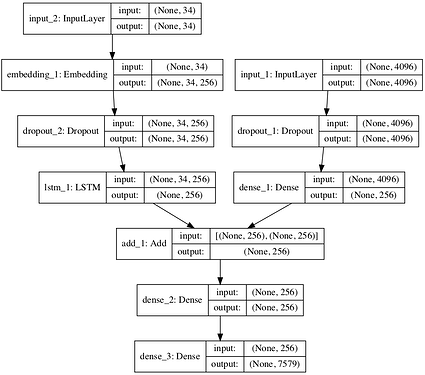
I was trying to implement this neural network in pytorch, but ran into some difficulties with an output of lstm layer.
The one thing i don’t understand why lstm layer on the model produced output in different shape from the input. There’s my implementation of this model. But the shape or rather dimension of the output stays the same as the input.
class Net(nn.Module):
def __init__(self):
super().__init__()
#LSTM
self.seq_embedding = nn.Embedding(51, 256)
self.seq_dropout = nn.Dropout(0.5)
self.seq_lstm = nn.LSTM(256, 256)
#Feature Extraction
self.feat_dropout = nn.Dropout(0.5)
self.feat_fc = nn.Linear(4096, 256)
#Decoder
self.fc = nn.Linear(256, 256)
self.fc2 = nn.Linear(256, 29045)
def forward(self, X1, X2):
out1 = self.seq_embedding(X1)
out1 = self.seq_dropout(out1)
out1, hidden = self.seq_lstm(out1)
out2 = self.feat_dropout(X2)
out2 = F.relu(self.feat_fc(out2))
out = torch.cat((out1, out2), 1)
out = self.fc(out)
out = self.fc2(out)
return F.softmax(out, dim=1)
my lstm dimmensions
Input: torch.Size([3656, 51, 256])
Output: torch.Size([3656, 51, 256])
Desired Output: torch.Size([3656, 256])
Is there any way to achieve the output from original model?