@Kaixhin Thanks for the link!
Sorry, I didn’t clarify enough. The MDN itself is not hard to implement as a concept (will, it depends on which distribution you will use for MDN). Since Alex Graves uses Gaussian Mixture Models for his MDN. With me, GMM required lots of work, and still quite unstable (however, I didn’t debug it further)
Hmm, I am not quite sure I agree (to be fair, I didn’t try this before). LSTM is not a magical tool at the end, it has a limited capacity. If you propagate the hidden state to the entire sequence (30,000), this hidden state isn’t a good enough representation to memorize such super long dependencies. Theoretically, it can model arbitrary length of sequences, but practically it doesn’t (this of course depends a lot on the data and the dimensionality that you have) (we ‘implicitly’ observe that even for image captioning, the LSTM can forget that it produced some words already. we are talking here about short sentences after all. To be fair, we didn’t explore this phenomena in depth. This is meta-analysis).
Also, if you use just 20 points, you are still taking advantage of the LSTM. The cell state is carried from one time step to another (from one point to another), so this is not equivalent to MLP at all.
A possible enhancement maybe to increase the sequence length.
Yes but he is doing 20 datapoints to make “one” prediction. An mlp would suffice for this. This data is just one float number per point in time series so 30,000 points does not constitute a lot of data. Maybe I’m bad explaining this so here is a link with a good explanation of using a stateful LSTM:
Thank you for the link, I will check it. I am interested to knew more about this concept (I don’t have this case in my work)


no problem. and to do something like this in pytorch you would just do something like:
output, (hx, cx) = model((input, (hx, cx))
and then in def forward:
def forward(self, inputs):
x, (hx, cx) = inputs
x = x.view(x.size(0), -1)
hx, cx = self.lstm(x, (hx, cx))
x = hx
return x, (hx, cx)
obviously a lot of other stuff in there for your desired outputs but thats the underlying basics to it
Ok now I’m confused 
Let me backtrack a bit to make sure I understand what you’re saying. You’re right because I’m reshaping my timeseries into a bunch of sequences of length q and use each one to predict the “next” observation. My input (and ground-truth) data is organized sequentially (timesteps 1-20 to predict 21, 2-21 to predict 22 etc). I then run nn.LSTM model with input size (20, batch_size, 1) to predict that “next” value. Now, you’re saying this is memoryless, and you also mentioned using nn.LSTMCell to create a stateful lstm in an earlier reply, are these two things related? This is where my confusion lies because I can’t see how my setup is preventing cell state to be carried from one time step to the next (I guess I just assumed nn.LSTM would automagically do this because… its an LSTM  ).
).
its not completely memoryless but it only useful for what happens from say i.e. 3-22 but has no recollection of what happened at 1 and 2 to help predict 23 and can only draw on information from 3-22. Have you seen this repo?:
It should help you
I say lstmcell cause its more intuitive cause its unrolled so output ready for next input as you can just pass cell state from rolled lstm to next lstm sequence as it should be only final part of that rolled up lstm
and yes u would think automatically work this way but for lots of other sequences this not advantageous i.e… translating a sentence, the order of words on previous sentence should have no bearing on how you would translate current sentence
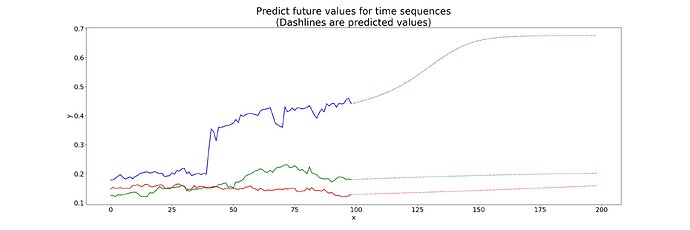
I did see the example a few weeks ago but its more useful now that I’ve been learning a bit more. If I replicate this method with the time series I posted in the OP I get the same results:
The prediction in the sine wave example is done like this:
for i, input_t in enumerate(input.chunk(input.size(1), dim=1)):
h_t, c_t = self.lstm1(input_t, (h_t, c_t))
h_t2, c_t2 = self.lstm2(c_t, (h_t2, c_t2))
outputs += [c_t2]
for i in range(future): # if we should predict the future
h_t, c_t = self.lstm1(c_t2, (h_t, c_t))
h_t2, c_t2 = self.lstm2(c_t, (h_t2, c_t2))
outputs += [c_t2]
I can see the final state c_t2 in the first loop is used as input for the first predicted pairs h_t, c_t in the second loop. The overall logic seems to be: take your last cell state and use it as input along with the second to last cell/hidden states, calculate new cell/hidden states, use these as inputs, repeat.
Why use h_t, c_t in the first cell instead of h_t2, c_t2 if these are the last cell/hidden states? I’m also confused about why there are two time steps involved. I’d appreciate if someone can break down the logic a bit more.
Ok so in that example you generate 100 sine waves consisting of 1000 step points.
To train he takes 97 of these sine waves and uses 999 step points to predict final 1000th values and trains on these using loss between output and target.(key here is that he predicts 999 outputs t2-t1000 not just the t1000 output)
As he finishes each epoch he test on the final 3 sine waves left over predicting 999 points but he also then uses last output c_t2 to do future loop to then make the next prediction but also because he also created his next (h_t, c,_t), ((h_t2, c_t2) in first iteration so has all he needs to propogate to next step and does for next 1000
“Why use h_t, c_t in the first cell instead of h_t2, c_t2 if these are the last cell/hidden states?”
He is using those for the future predicts remember in final test he does those first 999 loop those values still there to use and then does a 1000 future predictions
h_t, c_t For lstm1
h_t2, c_t2 for lstm2
Also, I think what can get confusing here is that this is during the forward the pass and the whole network is based off of manipulating this cell state in temporal order to get more accurate via training with Back Propagation Through Time
I do have to note sorry never noticed as pytorch example model it is quite lacking in the helpful comments with code category lol
Hi Alex, I am new to pytorch and also interested in time-series prediction.
I would like to ask some questions and appreciated you can share you idea.
In your training, did you reshape you input ?
Did you do loss = criterion(outputs, y) and how you match the dimension of output and y.
Thanks for breaking this down a bit, very helpful!
This was indeed a main source of confusion in my naive approach (I was predicting a single point t+1 from a size w window). Now I understand why you said earlier that the way I was setting up the problem was equivalent to a MLP (because there is no sequence for the LSTM to work with in the output).
Agreed. It’s fine since it’s an example and not a tutorial, but by god does it make things so much more confusing and difficult for beginners. On the upside it forces you to bleed through every single line of code, which is ok I guess if that’s your kink ![]()
Yes I did reshape my input. Check this post where there’s an ongoing discussion that might be helpful to you.
The way you reshape your input is key. I started with a target set of only a single timestep, which I believe doesn’t work well for LSTM as the target itself needs to be a sequence in order to apply recurrency.
Alex, thanks for the reply. I read the post you provided and I am confused about the shape.
“So if you divide a time series of length 10000 into chunks of length 50, your input tensor would be 50 (timesteps) by 200 (batch size) by 1 (features).”
"There are 200 batches in the dataset; each batch is 50x200x1."
For my understanding, so if we train only one batch? because 50*200=10000, 10000 is the total number of the data
so to sum up: if you want to train statefully a LSTM,
- don’t shuffle your batches
- use cell state and hidden state in the next forward pass to learn longterm dependencies
@chilango and @osm3000 did you try it out? if yes, what did you experience?
I didn’t try this (what you propose is called ‘stateful’ training if I remember correctly), although I expect it will work.
From my side, the conclusions (on generating sine wave) were:
- Sequence length is a super important factor.
- Just minimizing the prediction prediction (during the training) doesn’t reflect the expected the generation quality.
Since I first wrote about this experiments, we did more statistics on it in the recent period (sorry, I don’t have the graphs right now on this machine, I will upload them later). The results were super interesting (for me at least). I repeated this experiment for sequence lengths: 2, 5, 10, 20, 50, 100 timesteps (the cycle of the sine wave is 120 timesteps). For each sequence length, I trained 20 models, and I used each model to generate a sine-wave. I then evaluated each sine-wave (by hand) using a score from 0-3: 1 point for generating good amplitude, 1 for the frequency, and 1 for being centred around 0.
- The MSE for 2 & 5 is significantly worse than the rest.
- The MSE score for sequence lengths from 10 -> 100 is relatively the same
- I made a linear correlation between the sequence length/MSE and the generation quality:
~0.33 correlation factor between MSE and the generation quality
~0.95 correlation factor between sequence length and the generation quality
In short, I think there is more to this. We still have several hypotheses and experiments to do regarding this issue.
I went with @osm3000 suggestions of discretizing my time series and been playing around with a ton of models (although I think the discretization is a hack that I’d like to overcome at some point) . IIRC I got similar results in which after about 50 timesteps I noticed no difference in accuracy. I suspect however that the choice of timesteps (seq_len) should be more substantively driven, as in if you have weekly seasonality or daily seasonality you might want to tailor your sequence length to (multiples) of your period.
Another important factor in my experience is the resolution of the grid you use to discretize (if you decide to go with such an intermediary solution).
Haven’t worked on this project for a month or two now, but I remember that playing around with the depth of the network didn’t really have much of an impact beyond 2-3 layers. I’ve read elsewhere that such LSTMs can probably model most time series, but Graves (2014) uses an 8 layer network to model character RNNs.
Because I have very unbalanced classes I am using weighted learning which is easily implemented in pytorch.
HTH.
I am a bit surprised that discretization didn’t give you better results.
Is it possible you share the code?
Oh no by all means it did  Reframing into a classification problem makes a lot of sense because thats where the mature models are. What I meant is that eventually I’d like to understand MDN (GMM) enough to implement it in pytorch myself, so can move my predictions from discrete to continuous, so any pointers appreciated.
Reframing into a classification problem makes a lot of sense because thats where the mature models are. What I meant is that eventually I’d like to understand MDN (GMM) enough to implement it in pytorch myself, so can move my predictions from discrete to continuous, so any pointers appreciated.
Although this will sound strange, but the last research in Google (take a look at WaveNet Paper, https://arxiv.org/pdf/1609.03499.pdf , section 2.2. Also, check PixelRNN paper). I quote here:
However, van den Oord et al. (2016a) showed that a softmax distribution tends to work better, even when the data is implicitly continuous (as is the case for image pixel intensities or audio sample values). One of the reasons is that a categorical distribution is more flexible and can more easily model arbitrary distributions because it makes no assumptions about their shape.
Although these kind of statements should not be taken for granted (in my humble opinion), the very least to say - from my personal experience - is that life is much nicer with a softmax than a GMM. When I implemented AlexGraves model with GMM, it was quite unstable during the training (it diverges to inf, -inf, or nan, very easily). I tried several methods to control it, without success.