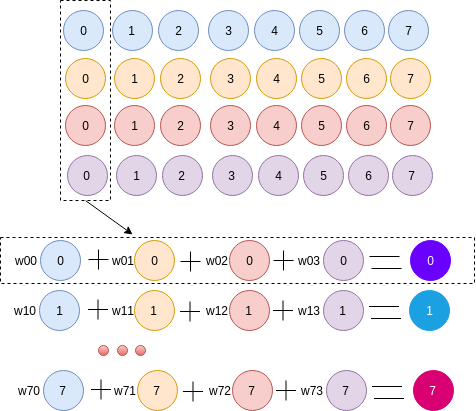
I have four vectors size of 1x8. I want to a new vector where each element row of the new vector is a linear combination of each element row in the four vectors as the figure. The expected output should be 1x8.
My solution is that using convolution to learn the weight. I convert the vector size of 1x8 to BxCxHxW, where W=1, and H=8, C is the number of vector. The combined vector has the size of 1x4x8x1. Now, we can use traditional conv2d with input size of 1x4x8x1 to produce the output size of 1x1x8x1. However, the output size (1x1x7x1) is mismatched with my expected. How should I solve it? How do you think my implementation of the problem. Is it kernel size = (4x1) or (1x1)?
import torch
import torch.nn as nn
class learn_vector(nn.Module):
def __init__(self, in_channels, out_channels):
super(learn_vector, self).__init__()
self.conv = nn.Conv2d(in_channels, 1, kernel_size=(in_channels, 1), stride=1, padding=(1,0), bias=True)
def forward(self, x):
print(x.size())
x = self.conv(x)
return x
batch_size, channel = 1, 8
input_vec1 = torch.randint(1, 100, (batch_size,channel)) # [1, 8]
input_vec2 = torch.randint(1, 100, (batch_size,channel)) # [1, 8]
input_vec3 = torch.randint(1, 100, (batch_size,channel)) # [1, 8]
input_vec4 = torch.randint(1, 100, (batch_size,channel)) # [1, 8]
# Extend vector from BxC --> BxC1xDxHxW where C=D
input_vec1 = input_vec1.view(batch_size, 1, channel, 1)
input_vec2 = input_vec2.view(batch_size, 1, channel, 1)
input_vec3 = input_vec3.view(batch_size, 1, channel, 1)
input_vec4 = input_vec4.view(batch_size, 1, channel, 1)
# Concat these vectors
input_concat = torch.cat([input_vec1, input_vec2, input_vec3, input_vec4], 1) #torch.Size([1, 4, 8, 1])
input_concat = input_concat.float()
# Feed to the network
learn_net = learn_vector(input_concat.size(1), 1)
out = learn_net(input_concat)
out = out.transpose (1,2) #torch.Size([1, 7, 1, 1])
print (out.size())