Hi all! I’m doing Time Series Prediction with the CNN-LSTM model, but I got overfitting condition. Here is my model code:
class LSTM(nn.Module):
def __init__(self, num_classes, input_size, hidden_size, num_layers, seq_length):
super(LSTM, self).__init__()
self.num_classes = num_classes
self.num_layers = num_layers
self.input_size = input_size
self.hidden_size = hidden_size
self.seq_length = seq_length
self.conv = nn.Conv1d(in_channels=num_classes, out_channels=num_classes,
kernel_size=(3,), padding=2,
padding_mode='replicate')
self.maxpool = nn.MaxPool1d(kernel_size=2, stride=1)
self.lstm = nn.LSTM(input_size=input_size, hidden_size=hidden_size,
num_layers=num_layers, batch_first=True, dropout=0.25)
self.batchnorm = nn.BatchNorm1d(hidden_size)
self.fc = nn.Linear(hidden_size, out_features=num_classes)
self.sig = nn.Sigmoid()
def forward(self, x):
x = x.permute(0,2,1)
conv_out = self.conv(x)
maxpool_out = self.maxpool(conv_out)
maxpool_out = maxpool_out.permute(0,2,1)
h_0 = torch.zeros(self.num_layers, x.size(0), self.hidden_size, requires_grad=True)
c_0 = torch.zeros(self.num_layers, x.size(0), self.hidden_size, requires_grad=True)
output_out, (h_out, c_out) = self.lstm(maxpool_out, (h_0, c_0))
out = output_out[:, -1, :]
batchnorm_output = self.batchnorm(out)
linear_output = self.fc(batchnorm_output)
sigmoid_output = self.sig(linear_output)
return sigmoid_output
And here is my train code:
seed = 23
torch.manual_seed(seed)
np.random.seed(seed)
num_epochs = 1000
learning_rate = 0.1
momentum = 0.9
input_size = 11
hidden_size = 32
num_layers = 2
num_classes = 11
model = LSTM(num_classes, input_size, hidden_size, num_layers, seq)
loss_func = nn.L1Loss()
print("-" * 70)
validator = True
optim_input = ""
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
scheduler = ReduceLROnPlateau(optimizer, mode='min', patience=75, verbose=True)
loss_list = []
epochs = []
time_per_epoch = []
# Pelatihan model
print("-" * 70)
print("Begin training with Convolutional LSTM layer (with Batch Normalization) and Adam optimizer\n")
model.train()
for epoch in range(num_epochs):
start = time.time()
outputs = model(train_x)
optimizer.zero_grad()
loss = loss_func(outputs, train_y)
loss.backward()
scheduler.step(loss)
optimizer.step()
stop = time.time()
epoch_time = stop - start
loss_list.append(loss.item())
epochs.append(epoch)
time_per_epoch.append(epoch_time)
print("Epoch: %d, MAE: %1.5f Time: %1.3f s" % (epoch, loss.item(), epoch_time))
print("-" * 70)
print(model)
print("-" * 70)
print("Training done in %1.5f seconds with last training MAE: %1.5f" % (sum(time_per_epoch), loss_list[-1]))
plt.figure(figsize=(10, 4))
plot_function(time_step=epochs, series=loss_list,
title='Training Loss Convolutional LSTM Layer with Batch Normalization\nOptimizer: Adam',
xlabel='Time', ylabel='Loss', label=["loss graph"])
For further information, my train set shape is [764,30,11] that consist of [batch_size,sequence_length,input_size] and my test set shape is [191,30,11]. My dataset is a normalized dataset with MinMaxScaler from scikit-learn. In training, I got an MAE of 0.01567, while in testing, I got an MAE of 0.16626, it’s a very huge difference between the normalized train and test.
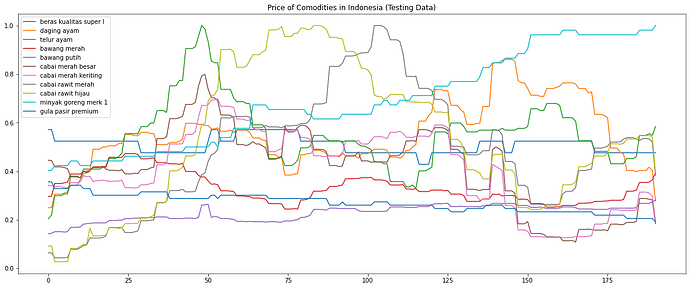
Here is the image of the test set:
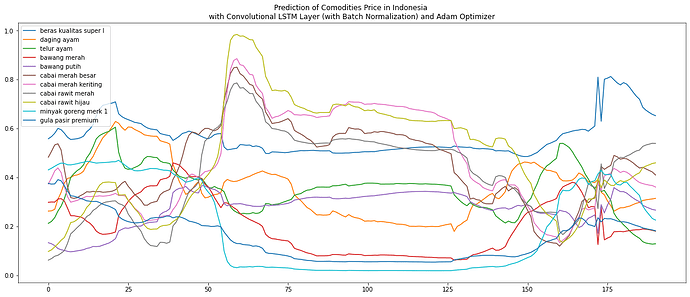
And here is the image of the prediction of the test set:
What am I supposed to do to reduce this overfitting condition?
Thank you in advance!