Tony-Y
July 27, 2019, 7:51am
21
The selection is based on the scores of frames. So, if the threshold is enough low, the number of selected frames could be larger than 1.
pcshih
July 27, 2019, 8:50am
22
“So, if the threshold is enough low” → I cannot understand this section.
The situation is probably like below.
Tony-Y
July 27, 2019, 9:54am
23
The decoder of FCSN consists of several temporal deconvolution operations which produces a vector of prediction scores with the same length as the input video. Each score indicates the likelihood of the corresponding frame being a key frame or non-key frame. Based on these scores, we select k key frames to form the predicted summary video.
Is the score described in the paper the probability that you use?
pcshih
July 27, 2019, 10:49am
24
The output (in the shape of [1,2,1,T]) of the FCSN is the score indicating whether to choose the frame or not. The scores actually are not limited to [0,1]. They can be any real number, because they do not add any activation function before FCSN output.
Thank you very much, Tony-Y.
Tony-Y
July 27, 2019, 11:05am
25
I think there are several approaches:
If Score(key) > Score(not) then it is a key frame.
If Score(key) > threshold then it is a key frame.
etc.
Tony-Y
July 27, 2019, 11:21am
26
By the way,
summary = x_select+h_select
summary should resemble x_select, but when summary equals to x_select, h_select is vanished.
pcshih
July 27, 2019, 12:33pm
27
I have done the first approach you mentioned in below
self.tanh_summary = nn.Tanh()
self.sigmoid = nn.Sigmoid()
def forward(self, x):
h = x
x_temp = x
h = self.FCSN(h)
values, indices = h.max(1, keepdim=True)
###old###
# # 0/1 vector, we only want key(indices=1) frame
# column_mask = (indices==1).view(-1).nonzero().view(-1).tolist()
# # if S_K doesn't select more than one element, then random select two element(for the sake of diversity loss)
# if len(column_mask)<2:
# print("S_K does not select anything, give a random mask with 2 elements")
# column_mask = random.sample(list(range(h.shape[3])), 2)
Thank you very much, Tony-Y.
I have reached the maximum number of replies a new user can create.issue
Tony-Y
July 27, 2019, 1:24pm
28
>>> import torch
>>> h = torch.randn(1,2,1,5, requires_grad=True)
>>> h.max(1, keepdim=True)
torch.return_types.max(
values=tensor([[[[-0.2387, 0.0180, 1.2485, -0.1497, -1.0964]]]],
grad_fn=<MaxBackward0>),
indices=tensor([[[[1, 0, 1, 0, 1]]]]))
The tensor indices also is non-differentiable.
pcshih
July 27, 2019, 2:09pm
29
I got the Basic badge 21 minutes ago.
I will try your suggestion in github .
But how to implement “index_mask = sigmoid( Score(key) -Score(not) )” in the output [1,2,1,T] tensor of the FCSN output?
Thank you very much, Tony-Y.
Tony-Y
July 27, 2019, 3:07pm
30
>>> import torch
>>> h = torch.randn(1,2,1,5)
>>> index_mask = torch.sigmoid( h[:,0] - h[:,1] )
>>> index_mask
tensor([[[0.7338, 0.7272, 0.6340, 0.0496, 0.2830]]])
pcshih
July 28, 2019, 2:31am
31
But the process “h[:,0], h[:,1]” are just like index_select, can they be diff. ?
Thank you very much, Tony-Y.
Tony-Y
July 28, 2019, 3:07am
32
It is a slice of the tensor.
pcshih
July 28, 2019, 5:20am
33
Although we get index_mask which is diff., how can we calculate reconstruction loss and diversity loss with this index_mask?
My problem is:
supposed index_mask=tensor([[[0.7338, -0.7272, 0.6340, -0.0496, 0.2830]]]), how can I convert index_mask to tensor([[[1, 0, 1, 0, 1]]])? and the process of conversion need to be diff.
Tony-Y
July 28, 2019, 6:53am
34
pcshih:
supposed index_mask=tensor([[[0.7338, -0.7272, 0.6340, -0.0496, 0.2830]]]),
No component of index_mask is negative because you use the sigmoid transformation of score.
Now, any component of index_mask is a real value in the range [0,1]. If you convert it to an integer value, there is no way to differentiate it.
pcshih
July 28, 2019, 7:27am
35
I have followed your idea but the loss is the same quite weird.
# if len(column_mask)<2:
# print("S_K does not select anything, give a random mask with 2 elements")
# column_mask = random.sample(list(range(h.shape[3])), 2)
# index = torch.tensor(column_mask, device=torch.device('cuda:0'))
# h_select = torch.index_select(h, 3, index)
# x_select = torch.index_select(x_temp, 3, index)
###old###
###new###
index_mask = self.sigmoid(h[:,1]-h[:,0]).view(1,1,1,-1)
#index_mask = (indices==1).type(torch.float32)
# if S_K doesn't select more than one element, then random select two element(for the sake of diversity loss)
# if (len(index_mask.view(-1).nonzero().view(-1).tolist()) < 2):
# print("S_K does not select anything, give a random mask with 2 elements")
# index_mask = torch.zeros([1,1,1,h.shape[3]], dtype=torch.float32, device=torch.device('cuda:0'))
# for idx in random.sample(list(range(h.shape[3])), 2):
# index_mask[:,:,:,idx] = 1.0
h_select = h*index_mask
x_select = x_temp*index_mask
###new reconstruct###
# diversity
S_K_summary_reshape = S_K_summary.view(S_K_summary.shape[1], S_K_summary.shape[3])
norm_div = torch.norm(S_K_summary_reshape, 2, 0, True)
S_K_summary_reshape = S_K_summary_reshape/norm_div
loss_matrix = S_K_summary_reshape.transpose(1, 0).mm(S_K_summary_reshape)
diversity_loss = loss_matrix.sum() - loss_matrix.trace()
#diversity_loss = diversity_loss/len(column_mask)/(len(column_mask)-1)
diversity_loss = diversity_loss/(index_mask.shape[-1])/(index_mask.shape[-1]-1)
S_K_total_loss = errS_K+reconstruct_loss+diversity_loss # for summe dataset beta=1
S_K_total_loss.backward()
# update
optimizerS_K.step()
##############
# update S_D #
##############
Thank you very much, Tony-Y.
Tony-Y
July 28, 2019, 7:49am
36
Your code is inconsistent with my proposed approach:
Tony-Y
July 28, 2019, 7:59am
37
By the way, can you reproduce the results of SUM-FCN ?
pcshih
July 28, 2019, 8:28am
38
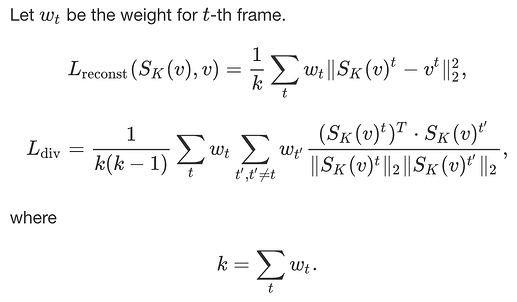
I revised my reconst. and div. loss
###new reconstruct###
reconstruct_loss = torch.sum((S_K_summary-vd)**2 * index_mask) / torch.sum(index_mask)
###new reconstruct###
# diversity
S_K_summary = index_mask*S_K_summary
S_K_summary_reshape = S_K_summary.view(S_K_summary.shape[1], S_K_summary.shape[3])
norm_div = torch.norm(S_K_summary_reshape, 2, 0, True)
S_K_summary_reshape = S_K_summary_reshape/norm_div
loss_matrix = S_K_summary_reshape.transpose(1, 0).mm(S_K_summary_reshape)
diversity_loss = loss_matrix.sum() - loss_matrix.trace()
#diversity_loss = diversity_loss/len(column_mask)/(len(column_mask)-1)
diversity_loss = diversity_loss/(torch.sum(index_mask))/(torch.sum(index_mask)-1)
Reproduction of SUM-FCN is FCSN.py file.
Thank you very much, Tony-Y.
Tony-Y
July 28, 2019, 8:33am
39
Is your results of SUM-FCN consistent with the original results?