hello everyone ,
I’m coding the dropout as a variational inference as [ Gal et al 2016 ] explain it
I have a problem regarding “BCE With Logits Loss”
My simple neural net is
def __init__(self, hidden_size, input_size=1, dropout_p=0.25 ):
super().__init__()
self.dropout_p = dropout_p
self.hidden1 = nn.Linear(input_size,hidden_size)
self.relu = nn.ReLU()
self.output = nn.Linear(hidden_size,1)
def forward(self, x):
x = self.relu(self.hidden1(x))
x = F.dropout(x,p=self.dropout_p)
x = self.output(x)
x = torch.sigmoid(x)
return x
bnn = MLP(hidden_size=20,input_size=2)
bnn.train()
criterion = nn.BCELoss()
optimizer_bnn = torch.optim.SGD(bnn.parameters(), lr=0.01, momentum=0.90, nesterov=True, weight_decay=1e-6)
my training loop
torch.manual_seed(7)
fig, ax = plt.subplots(figsize=(7,7))
num_epochs=200
# To Do here
######################### classical Gradient training loop #################################
for epoch in range(num_epochs):
for (X,Y) in train_dataloader:
# Forward pass
outputs = bnn(X.float())
loss = criterion(outputs.float().reshape(-1), Y.float())
# Backward and optimize
optimizer_bnn.zero_grad()
loss.backward()
optimizer_bnn.step()
# For plotting and showing learning process at each epoch, uncomment line below
if (epoch+1)%10==0:
plot_decision_boundary( bnn, X, Y, epoch, ((outputs.squeeze()>=0.5) == Y).float().mean(),
nbh=4, model_type='mcdropout')
##############################################################################################
print('Finished Training')
My plot function
# Useful function: plot and show learning process in classification
def plot_decision_boundary(model, X, Y, epoch, accuracy, model_type='classic', samples=100, nbh=2, cmap='RdBu'):
h = 0.02*nbh
x_min, x_max = X[:,0].min() - 10*h, X[:,0].max() + 10*h
y_min, y_max = X[:,1].min() - 10*h, X[:,1].max() + 10*h
xx, yy = np.meshgrid(np.arange(x_min*2, x_max*2, h),
np.arange(y_min*2, y_max*2, h))
test_tensor = torch.from_numpy(np.c_[xx.ravel(), yy.ravel()]).float()
if model_type=='classic':
model.eval()
pred = model(test_tensor)
elif model_type=='svi':
pred = model.forward(test_tensor, n_samples=samples).mean(0)
elif model_type=='mcdropout':
model.eval()
model.training = True
outputs = torch.zeros(samples, test_tensor.shape[0], 1)
for i in range(samples):
outputs[i] = model(test_tensor)
pred = outputs.mean(0).squeeze()
Z = pred.reshape(xx.shape).detach().numpy()
plt.cla()
ax.set_title('Classification Analysis')
ax.contourf(xx, yy, Z, cmap=cmap, alpha=0.25)
ax.contour(xx, yy, Z, colors='k', linestyles=':', linewidths=0.7)
ax.scatter(X[:,0], X[:,1], c=Y, cmap='Paired_r', edgecolors='k');
ax.text(-4, -7, f'Epoch = {epoch+1}, Accuracy = {accuracy:.2%}', fontdict={'size': 12, 'fontweight': 'bold'})
display.display(plt.gcf())
display.clear_output(wait=True)
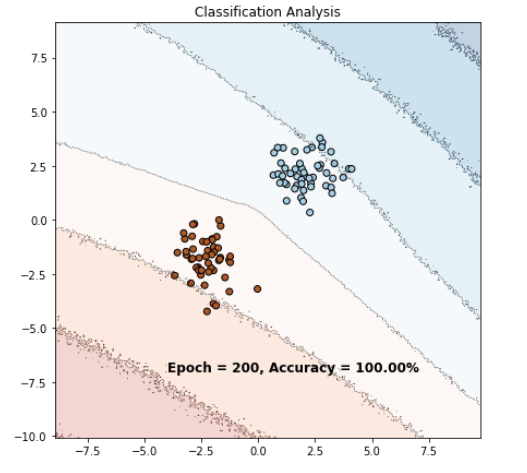
My problem is if I keep the sigmoid in the forward and use BCELoss I get this final result
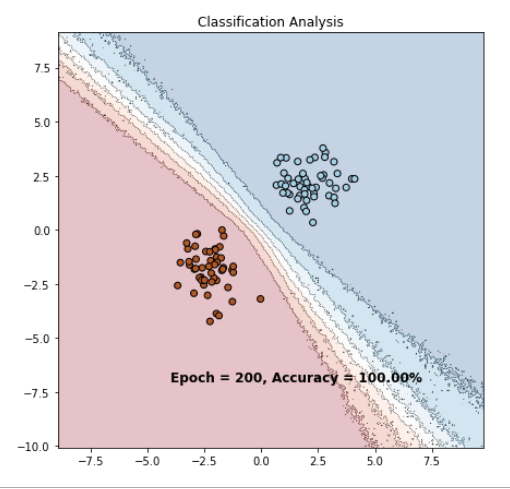
However if I change it without a sigmoid function and use BCEwithLogits I get this result