Hi Tom,
Wow - that is a perfect summary - it took me a nearly a full days reading to come to the same level of understand.
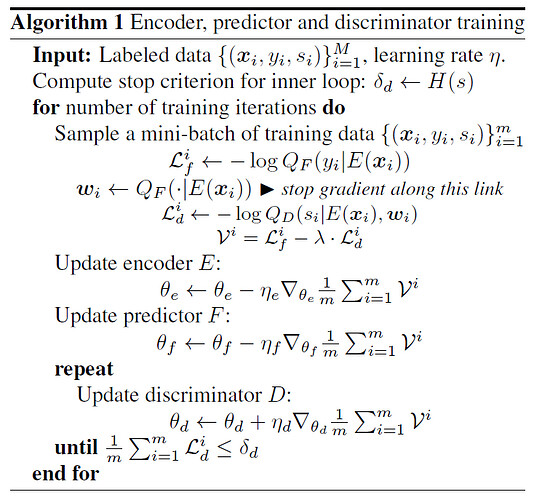
Indeed in this case the purpose of the discriminator network D, is to remove conditional dependencies on the sources (i.e. sleeping subject + measurement environment).
Perhaps you meant e = e(sample) rather than e=e(source)? As the encoder, is fed in the sequence of spectrograms X = [x_1,x_2,…,x_t] in Omega_x, up to some time t, in the notation of the start of section 3. Model.
At the moment my use/test case is pretty much the same as the original paper - but due to the generality of this domain adaptation/source conditionally independent, type of setup I guess it could be applied to many use cases in general time-series classification and prediction?
The minimum requirement’s would be as you clearly summarised, (a time-series sample X, source ID s, instantaneous event labels y). So I guess there could be applications to claim modelling/event classification & prediction in insurance - the solid proof/performance guarantee of the paper makes it a good candidate?
My guess is due to the generality of this setup it could eventually be extended to extra modalities, by simply training more independent encoders say a, b, c, on each of them, and then concatenating the outputs of these encoders - and projecting this vector, e.g.
E_canonical (X_a, X_b, X_c) = concatenate [ E_a(X_a), E_b(X_b), E_c(X_c)]
onto a latent embedding/manifold. The projection from this canonical space, could then be fed into the predictor and discriminator networks as before. Some notion of how perhaps to attempt this is given here - Conditional generation of multi-modal data using constrained embedding space mapping. It’s just an idea at the moment though. Perhaps it could again be very applicable in actuarial science, maybe as an alternative to classical Gaussian/Levy/Poisson Process models - given enough data it should hopefully pick up notions of correlations in both time and across modalities?
I’m not sure about the application to style fingerprinting, so maybe we could just try it and see?
I have some test data which is similar to that used in this paper. The difference is, the available data uses accelerometer recording’s (rather than radio frequency modulations) of sleep study participants, and it also has their “gold standard” labels from polysomnography (PSG).
https://es.informatik.uni-freiburg.de/datasets/ichi2014
So converting this 3 channel accelerometer data to sequences of spectrograms is reasonably simple to do. After that the papers encoder architecture could be applied, and it’s algorithm should be reproducible without any modifications - I hope 
Hopefully in a few months time I should have access to some SOTA clinical grade wearable sensors, which record ECG, electrodermal activity, respiration rate/depth - generally multi-modal streaming bio-markers of sympathetic nervous system and cardio-respiratory activities - similar to the Verily Study Watch.
I’m guessing something similar to that will be the new standard recording instrument in health insurance studies, in a few years time?
Great to be chatting with you again 
Best regards,
Ajay
![]()
![]()