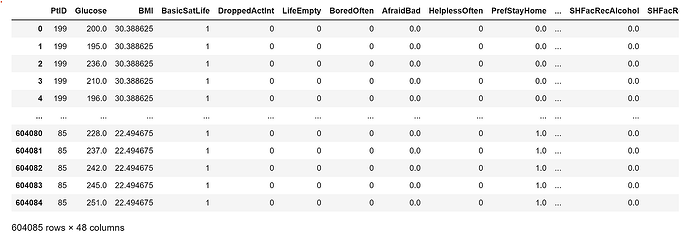
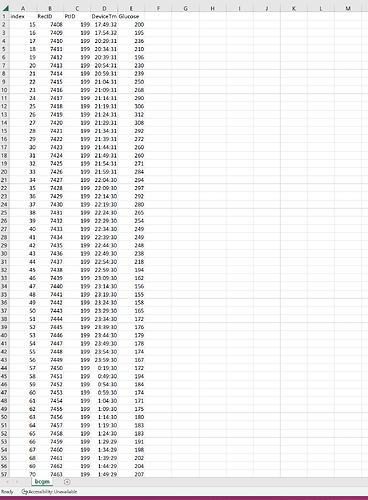
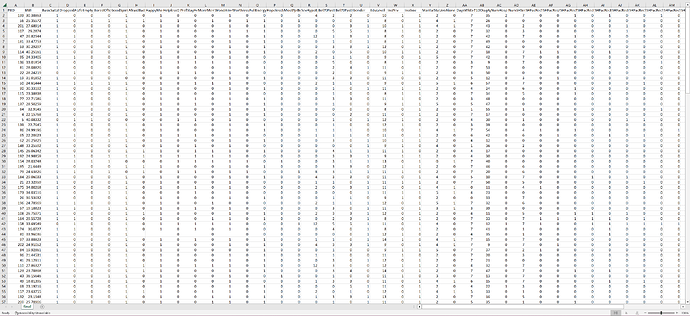
For starters, the DeviceTm has time gaps. So you will either need a positional encoder. Or you can fill time gaps with padding. See one of my earlier posts in this thread for a link for how to set up a position encoder.
Then you need to set the __len__ in the Dataset. In this case, you can use np.unique() targetted on the correct row for PtID. That will give you an array of all unique patient IDs. And then get the length of that array to insert in __len__. But store the unique values array because you can use that in the __getitem__ function.
The above assumes that each Patient ID has the same number of data points.
What is still not clear to me is whether each PtID has an equal number of glucose readings. Now let’s assume they do not. In that case, you can set an arbitrary length for the input sequence, say 40 (i.e. the last 2 hours) and then prepare your data samples such that each consecutive 2 hours and 30 minutes of data, where there is a glucose reading at the final time(i.e. at 2h and 30m), comprises a sample. Two hours for model input data and the final time for your target.
So the __len__ value would also need to be defined differently.
In the above method, you’d need to initially fill in all time gaps with a zero reading. This is how you can do that with Pandas, except you can put 0 instead of NaN.
Second, you’d need to count the number of non-zero targets that have at least 45 data points prior. I.e.
total_samples = 0
PtID_idx = 2 # 2 if id is in column C, otherwise adjust
Glucose_idx = 4
for id in unique_patients:
patient_glucose = x[:,x[PtID_idx,:]==id] # filter all of the values that correspond with that patient id
#can fill patient_glucose gaps with zeros here, with the above linked method; just get it back into numpy before continuing
try:
patient_glucose = patient_glucose[:,45:] #remove first 45 values for that patient
except:
continue
patient_glucose = patient_glucose[:,patient_glucose[Glucose_idx,:]==0] # remove the gaps, remaining values will be added to total sample count, since targets should be non-zero
total_samples += patient_glucose.shape[1] # add values to targets
You may need to add an additional condition for excluding gaps 45 or larger(or possibly even 20 and larger).
From there, you can use similar conditioning arguments in __getdata__ to acquire your samples from both data tables.
The dataset is small enough that you could preprocess it all in advance and store it in memory.