Hello,
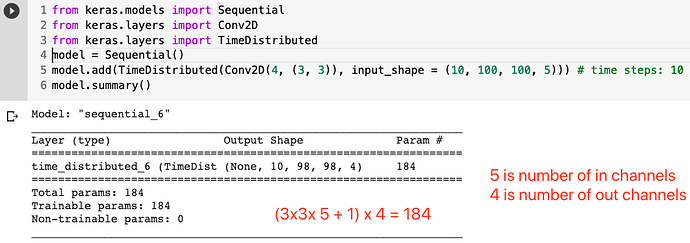
I am implementing a paper’s architecture that does Time distributed CNN over the input. For the sake of clarification and with the input in the form of (batch_size, time_steps, channels, H, W):
let’s say the input is (32, 100, 1, 128, 128) and after applying the convolution with 16 kernels I get (32, 100, 16, 64, 64).
after reading through the forum, I used the trick of multiplying the dimensions batch_size and time_steps, and then reshape when done with convolution.
Okay, I tried this, and the shapes are good, and code runs well. However, when I checked the number of parameters in my model, it was less than that in the paper. And I believe the reason is, it is using the same kernel weights over the whole batch and time_steps.
Which means blending the dimensions isn’t doing actually timeDistributed convolution. Any insights to solve the problem ?
Thank you