for i in range(realization):
condiMI=
ground_truth=
result_ITE=
for thresh in thresh_vec:
rho_data = data_generation(rho, thresh, size)
modelP = Class_Net(input_size = 3, hidden_size = 130, std = 0.08)
modelQ = Class_Net(input_size = 2, hidden_size = 100, std = 0.02)
optimizerP = torch.optim.Adam(modelP.parameters(), lr=1e-3)
optimizerQ = torch.optim.Adam(modelQ.parameters(), lr=1e-3)
# conditional mi
mi_p = mi(rho_data, size, train_size, modelP, optimizerP, rep, tau, input_size=3)[0]
mi_q = mi(rho_data, size, train_size, modelQ, optimizerQ, rep, tau, input_size=2)[0]
condi_mi = mi_p - mi_q
condiMI.append(condi_mi*1.4427)
# ground truth
p=scipy.stats.norm(0, 1).cdf(thresh)
ground_value=-(1-p)*0.5*np.log(1-rho*rho)*1.4427
ground_truth.append(ground_value)
print("Conditional TE", condiMI)
print("ground_truth", ground_truth)
# ite
vae_net=VAE()
vae_net.to(device)
diff_ite=[]
modelA = Class_Net(input_size=3, hidden_size=130, std=0.08)
modelB = Class_Net(input_size=2, hidden_size=100, std=0.02)
optimizerA = torch.optim.Adam(modelA.parameters(), lr=1e-3)
optimizerB = torch.optim.Adam(modelB.parameters(), lr=1e-3)
for j in range(repo):
vae_data, Jacobian_joint=data_gen_zbar(rho_data, size, vae_net)
#jacobian matrix is equal to Jacobian_joint together with Jacobian_joint by mar_index
miA, gradsA, indexA, t_indexA, m_indexA = mi(vae_data, size, train_size,
modelA, optimizerA, repi,
tau,input_size=3)
J_reorderA = torch.index_select(Jacobian_joint, 1, torch.LongTensor(m_indexA).cuda())
J_Am1 = torch.index_select(J_reorderA, 1, torch.LongTensor(t_indexA).cuda())
J_Aj1 = torch.index_select(Jacobian_joint, 1, torch.LongTensor(t_indexA).cuda())
Jacobian_jm = torch.cat ((J_Aj1, J_Am1), 1)
Jacobian_A = torch.index_select(Jacobian_jm, 1, torch.LongTensor(indexA).cuda())
miB, gradsB, indexB, t_indexB, m_indexB = mi(vae_data, size, train_size,
modelB, optimizerB, repi,
tau, input_size=2)
J_reorderB = torch.index_select(Jacobian_joint, 1, torch.LongTensor(m_indexB).cuda())
J_Bm1 = torch.index_select(J_reorderB, 1, torch.LongTensor(t_indexB).cuda())
J_Bj1 = torch.index_select(Jacobian_joint, 1, torch.LongTensor(t_indexB).cuda())
Jacobian_jmB = torch.cat ((J_Bj1, J_Bm1), 1)
Jacobian_B = torch.index_select(Jacobian_jmB, 1, torch.LongTensor(indexB).cuda())
grads_j_A=gradsA[:,-1].reshape(-1,1)
grads_j_B=gradsB[:,-1].reshape(-1,1)
#calculate gradient
#calculate the gradient wrt the weights of network vae
grads_A=torch.mm(torch.t(grads_j_A),torch.t(Jacobian_A))
grads_B=torch.mm(torch.t(grads_j_B),torch.t(Jacobian_B))
diff_grads=grads_A-grads_B
with torch.no_grad():
vae_net.fc1.weight -=alpha*torch.t(diff_grads)
diff_ite.append(miA-miB)
result_ITE.append(ma(diff_ite)[-1]*1.4427)
print("result_ITE", result_ITE)
total_te[i,:] = condiMI
total_ite[i,:] = result_ITE
final_result_STE=[a_i - b_i for a_i, b_i in zip(condiMI, result_ITE)]
total_ste[i,:]=final_result_STE
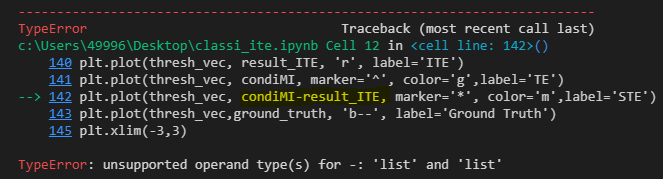
plt.plot(thresh_vec, condiMI-result_ITE, marker=‘*’, color=‘m’,label=‘STE’)
Hi guys, i have met this problem here. In this code, occurs type error:
plt.plot(thresh_vec, condiMI-result_ITE, marker=‘*’, color=‘m’,label=‘STE’)
TypeError: unsupported operand type(s) for -: ‘list’ and ‘list’