Hi
I’m trying to implement a U-Net in PyTorch. I have been following this article: https://towardsdatascience.com/u-net-b229b32b4a71
I have created binary masks in MatLab using poly2mask for about 650 images. The binary masks are in BMP format and consists of 256x256 pixels with values of either 0 or 255 (i.e. black background and white object).
I load the training images with corresponding masks into tensors:
orgdir = "\image_unet\org"
maskdir = "\image_unet\mask
for k in range(0,n_train):
# Load Image
number = "_{}.bmp".format(k+1)
orgname = "org"+number
maskname = "mask"+number
im = io.imread(os.path.join(orgdir,orgname))
im = ts.Tensor(im)
img_train[k,:,:,:]=im
mask = io.imread(os.path.join(maskdir,maskname))
mask = ts.Tensor(mask)
mask = mask / 255
label_train[k,:,:,:] = mask
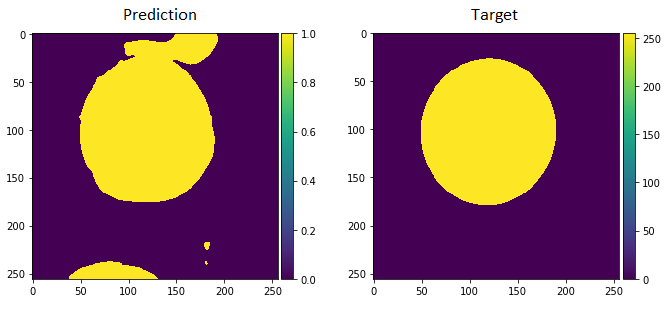
I am able to get an output from the U-Net I have made from the article:
class UNet(nn.Module):
def encoding_block(self, in_channels, out_channels, kernel_size=3):
block = nn.Sequential(
nn.Conv2d(in_channels=in_channels, out_channels=out_channels, kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(out_channels),
nn.Conv2d(in_channels=out_channels, out_channels=out_channels,kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(out_channels),
)
return block
def decoding_block(self, in_channels, mid_channels, out_channels, kernel_size=3):
block = nn.Sequential(
nn.Conv2d(in_channels=in_channels, out_channels=mid_channels, kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(mid_channels),
nn.Conv2d(in_channels=mid_channels, out_channels=mid_channels, kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(mid_channels),
nn.ConvTranspose2d(in_channels=mid_channels, out_channels=out_channels, kernel_size=kernel_size,stride=2, padding=1,output_padding=1),
)
return block
def final_block(self, in_channels, mid_channels, out_channels, kernel_size=3):
block = nn.Sequential(
nn.Conv2d(in_channels=in_channels, out_channels=mid_channels, kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(mid_channels),
nn.Conv2d(in_channels=mid_channels, out_channels=mid_channels, kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(mid_channels),
nn.Conv2d(in_channels=mid_channels,out_channels=out_channels,kernel_size=kernel_size,padding=1),
nn.ReLU(),
nn.BatchNorm2d(out_channels),
)
return block
def __init__(self, in_channels, out_channels):
super(UNet, self).__init__()
# Encoding:
self.encode_1 = self.encoding_block(in_channels=in_channels,out_channels=64)
self.maxpool_1 = nn.MaxPool2d(kernel_size=2)
self.encode_2 = self.encoding_block(in_channels=64,out_channels=128)
self.maxpool_2 = nn.MaxPool2d(kernel_size=2)
self.encode_3 = self.encoding_block(in_channels=128,out_channels=256)
self.maxpool_3 = nn.MaxPool2d(kernel_size=2,ceil_mode=True)
# From Encoding to Decoding:
self.enc2dec = nn.Sequential(
nn.Conv2d(in_channels=256,out_channels=512,kernel_size=3,padding=1),
nn.ReLU(),
nn.BatchNorm2d(512),
nn.Conv2d(in_channels=512,out_channels=512,kernel_size=3,padding=1),
nn.ReLU(),
nn.BatchNorm2d(512),
nn.ConvTranspose2d(in_channels=512,out_channels=256,kernel_size=3, stride=2, padding=1,output_padding=1)
)
# Decoding:
self.decode_3 = self.decoding_block(in_channels=512,mid_channels=256,out_channels=128)
self.decode_2 = self.decoding_block(in_channels=256,mid_channels=128,out_channels=64)
self.final = self.final_block(in_channels=128,mid_channels=64,out_channels=out_channels)
def crop_and_concatenate(self, upsample, encoded_block, crop=False):
#print("Size upsamle = {}".format(upsample.size()))
#print("Size encoded_block = {}".format(encoded_block.size()))
if crop:
c = (encoded_block.size()[2] - upsample.size()[2]) // 2
encoded_block = F.pad(encoded_block, (-c,-c,-c,-c))
return ts.cat((upsample,encoded_block),1)
def forward(self, x):
# Encoding
encode_block_1 = self.encode_1(x)
encode_pool_1 = self.maxpool_1(encode_block_1)
encode_block_2 = self.encode_2(encode_pool_1)
encode_pool_2 = self.maxpool_2(encode_block_2)
encode_block_3 = self.encode_3(encode_pool_2)
encode_pool_3 = self.maxpool_3(encode_block_3)
# Encoding to decoding
bottleneck = self.enc2dec(encode_pool_3)
# Decoding
decode_block_3 = self.crop_and_concatenate(bottleneck, encode_block_3, crop=True)
concatenation_layer_2 = self.decode_3(decode_block_3)
decode_block_2 = self.crop_and_concatenate(concatenation_layer_2, encode_block_2, crop=True)
concatenation_layer_1 = self.decode_2(decode_block_2)
decode_block_1 = self.crop_and_concatenate(concatenation_layer_1, encode_block_1, crop= True)
final_block = self.final(decode_block_1)
return ts.sigmoid(final_block)
I am using BCELoss() and this is where the problems begin. The loss increases for each epoch. It begins from about 0.4 and then increases steadily with about 1.0 for each epoch. My training function looks like:
def train(network, img, target):
criterion = nn.BCELoss()
#optimizer = ts.optim.SGD(network.parameters(),lr=0.01)
optimizer = ts.optim.Adam(network.parameters())
running_loss = 0
epoch_loss = 0
for epoch in range(100):
print("Epoch {} out of {}".format(epoch+1,10))
for i, im in enumerate(img):
# status
if (i % 50) == 0:
print("Image {} out of {}".format(i,img.shape[0]))
# get input
image, truth = im.view(1,1,256,256), target[i].view(1,2,256,256)
# variable wrap
image, truth = ts.autograd.Variable(image), ts.autograd.Variable(truth)
# forwardpass, backprop, optimization step
prediction = network(image)
#print(prediction.shape)
#prediction = prediction.view(-1)
#prediction = F.softmax(prediction,dim=0)
# zero gradients
optimizer.zero_grad()
#truth = truth.view(-1)
loss = criterion(prediction, truth)
running_loss += loss.item()
loss.backward()
optimizer.step()
epoch_loss = running_loss / (i+1)
print('Loss for epoch {} was {}'.format(epoch+1,epoch_loss))
return network
As I am very new to deep learning I am really in doubt where I go wrong and how I can fix it. My idea was: Input an original image, then output a single feature map 256x256x1 and compute the binary cross entropy loss with the mask corresponding to the input image also with dimension 256x256x1, but this idea appears to be wrong.
In the U-Net article, they output two feature maps where one is for background and one is for object, I think, so my preprocessing might be misguided.
I really hope someone can point me to where I go wrong: Is it the image-mask preprocessing? Is it the code? Have I misunderstood the loss computation?
I would really appreciate any inputs.
Thanks in advance,
Simon