Hey, everyone!
I am trying to do a segmentation on T1 and FLAIR for multiple sclerosis. Currently i am trying to load each patient to dataloader, where batchsize is number of patients. The images are concatinated and had the shape (Depth, flair+T1, height, width), when they come out from the dataloader, i get this shape (batchsize, Depth, flair+T1, height, width). So what I am trying to do is to send chunks of continous slices of the brain to a unet, until the network has seen the entire brain, and then bsckpropagete (if this method is correct). What i am stumbling on is memory error on the gpu. It crashes for any size of batchsizes. I think the problem is the gradients accumulated, but i dont know how to fix that…
My setup:
Amd ryzen 7
16gb ram
Nvidia RTX 3070 gb
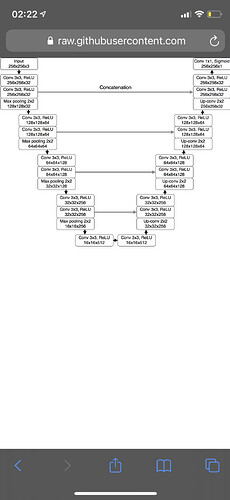
My unet:
And my train code:
import argparse
import os
import numpy as np
import torch
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from tqdm import tqdm
import argparse
from loss import DiceLoss, TverskyLoss
from torchvision import transforms, utils
from unet import UNet
from preproccess import normalize, ToTensor, check_axis,add_depth_channel, ConcatenateVolumes
from NiftiLoader import MSdataset
import matplotlib.pyplot as plt
class main:
def __init__(self, args):
self.args = args
self.train_loader = None
self.in_channel = None
self.out_channel = None
def _config_dataloader(self):
print("Starting configuration of the dataset")
print("Collecting validation and training set")
validation_mode = "/validation/"
training_mode = "/train/"
train_dataset = MSdataset(self.args.path + training_mode, composed_transforms = [
check_axis(),
add_depth_channel(depth = self.args.depth),
ToTensor(),
normalize(z_norm = True)]
)
validation_dataset = MSdataset(self.args.path + validation_mode, composed_transforms = [
check_axis(),
add_depth_channel(depth = self.args.depth),
ToTensor(),
normalize(z_norm = True)]
)
train_loader = DataLoader(train_dataset,
self.args.patient_batch_size,
self.args.shuffle)
validation_loader = DataLoader(validation_dataset,
self.args.patient_batch_size,
self.args.shuffle)
#for idx, smpls in train_loader:
# print(idx, smpls)
print("Data collected. Returning dataloaders for training and validation set")
return train_loader, validation_loader
def __call__(self, is_train = True):
train_loader, validation_loader= self._config_dataloader()
complete_data = {"train": train_loader, "validation": validation_loader }
if torch.cuda.is_available():
print("Using CUDA")
device = torch.device("cuda:0")
else:
print("Using CPU")
device = torch.device(‘cpu’)
unet = UNet(in_channels=2, out_channels=1, init_features=32)#,
unet.to(device, dtype=torch.float32)
optimizer = optim.Adam(unet.parameters(), lr=self.args.lr)
#dsc_loss = DiceLoss()
Tloss = TverskyLoss()
loss_train = []
loss_validation = []
print("Starting training process. Please wait..")
sub_batch_size = 8
for current_epoch in tqdm(range(self.args.epoch),total= self.args.epoch):
print(f"current epoch {current_epoch}/{self.args.epoch}")
for phase in ["train", "validation"]:
if phase == "train":
unet.train()
if phase == "validation":
unet.eval()
for i, data_set_batch in enumerate(complete_data[phase]):
X, mask = data_set_batch["volume"], data_set_batch["mask"] #<--
#X, mask = (X.to(device)).float(), mask.to(device)
N = X.shape[1]//sub_batch_size
#patient,D,C,H,W = X.shape #dette er shapen fra dataloaderen.
#mask =mask.reshape((patient*D,H,W)) # <--dette er en pasient
#X = X.reshape((patient*D,C,H,W)) #<--samme pasient
optimizer.grad_zero()
with torch.set_grad_enabled(phase == "train"):
for current_patient in range(X.shape[0]):
train_loss_depths = 0
validation_loss_depths = 0
optimizer.zero_grad()
for sub_batches in tqdm(range(0,X.shape[1]- sub_batch_size, sub_batch_size)):
mask_input = mask[current_patient, sub_batches: sub_batches + sub_batch_size,:,:]
mask_input = mask_input.to(device, dtype = torch.float32)
small_batch_input = X[current_patient, sub_batches: sub_batches + sub_batch_size,:,:,:]
small_batch_input = small_batch_input.to(device, dtype = torch.float32)
predicted = unet(small_batch_input)
predicted = predicted.squeeze(1)
loss = Tloss(predicted, mask_input)
if phase == "train":
train_loss_depths = (train_loss_depths + loss)
if phase == "validation":
validation_loss_depths = (validation_loss_depths + loss)
if phase == "train":
train_loss_depths = (train_loss_depths)/N
print("running loss train:", train_loss_depths, "\n")
loss_train.append(train_loss_depths.item())
train_loss_depths.backward()
optimizer.step()
if phase == "validation":
loss_validation.append(validation_loss_depths.item()/N)
print("running validation loss", validation_loss_depths.item()/N, "\n")
This is just a small and unfinished example of my train code.