I have a UNET model loaded from the SMP library to semantically segment the DDSM database:
class LesionSegmentation(nn.Module):
def __init__(self):
super(LesionSegmentation, self).__init__()
self.device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
self.model = smp.Unet(
encoder_name="resnet34",
encoder_weights="imagenet",
in_channels=1,
classes=1,
activation='sigmoid'
)
def forward(self, x):
# Perform forward pass through model
mask = self.model(x)
return mask
I am using the Dice Loss:
def dice_score(y_true, y_pred):
return (2 * (y_true * y_pred).sum() + 1e-15) / (y_true.sum() + y_pred.sum() + 1e-15)
Images and masks are both B, 1, 224, 224
So far the Dice Loss has proven to be the best in terms of accurately representing my model because other losses, e.g., BCE will be really low (0.005) with a mean IOU of 0.1
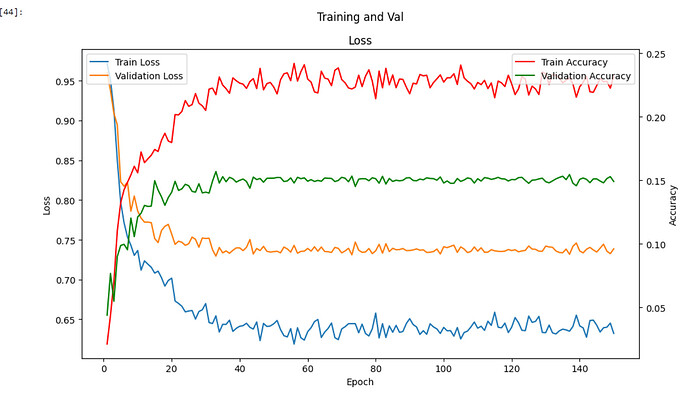
My problem is, no matter what I edit, e.g., loss fun, encoder, model etc. My results are the same:
The problem is, I don’t know where to go from here. I’ve trialled all sorts of loss functions because I thought that maybe there is a class imbalance for my dataset in terms of lesion/nonlesion ratio
This is the dataset I’m using: The Complete Mini-DDSM | Kaggle