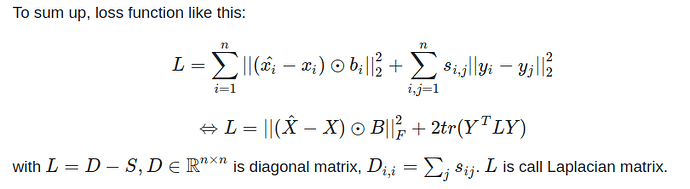Hi, I am trying to implement SDNE, a algorithm uses deep auto encoder to map a graph to latent representation d dimension.
The idea is kind of simple, SDNE uses the adjacency matrix as input and output is embedding layer (also node embedding of graph). The loss function tries to minimize the reconstruction layer and input layer which defined $\hat{X}$ and $X$. To keeping node embedding Y (embedding layer) is close together like the original node in graph input, loss function continue minimizes $||y_i - y_j||*s_{ij}$ with $s_{ij}$ is weight of node $v_i$ and $v_j$, if not link $s_{ij} = 0$.
When I implement code in PyTorch, I got negative loss value although the loss function does not negative value.
This is my code:
def _compute_loss(self, x, x_hat, y, L):
def loss_1st(Y, L):
Y_ = torch.matmul(torch.transpose(Y, 0, 1), L)
return 2 * torch.trace(torch.matmul(Y_, Y))
def loss_2nd(X_hat, X, beta):
B = torch.ones_like(X).to(device)
B[X != 0] = beta
return torch.sum(torch.square((X_hat - X) * B))
batch_size = x.shape[0]
# TODO: check if divide batch_size
loss_1 = loss_1st(y, L)
loss_2 = loss_2nd(x_hat, x, self.beta)
loss = loss_2 + self.alpha * loss_1
return loss
I trained on all batch dataset which means batch_size = len(Adjancency_matrix) = len(A).
Actually, I’m not sure about this algorithm can train on batch_size = 64, 128,… because I can not calcualte L on mini batch and mini batch also can not represent for the whole graph.
So, can you see in my code what reason why my loss function is negative?
Thanks!