Ahhh that’s great, it’s so simple and clean! ![]() Thank you for having a look! Out of curiosity is there a way to allow the gradient to flow back to the indices as well? In my case these are actually something that needs learning alongside the scattered values…it’s a strange network!
Thank you for having a look! Out of curiosity is there a way to allow the gradient to flow back to the indices as well? In my case these are actually something that needs learning alongside the scattered values…it’s a strange network!
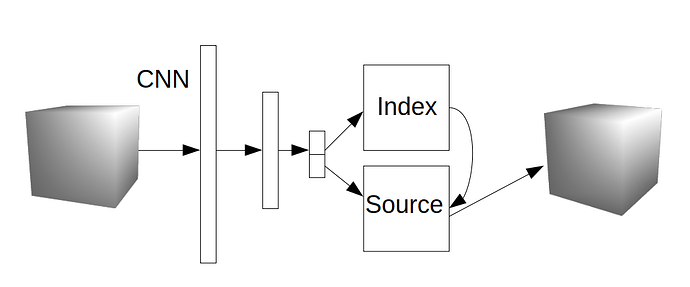
Think, something like this, where Index is i in the above visualisation, Source is v, and the output cube is z.
Here, I think because the index is something that is learned through the CNN part of the network there should be some way to allow for the gradients to flow back through the Index tensor. (See related post for this argument)
Edit: Perhaps using the out-of-place operation torch.Tensor.scatter() solves this?