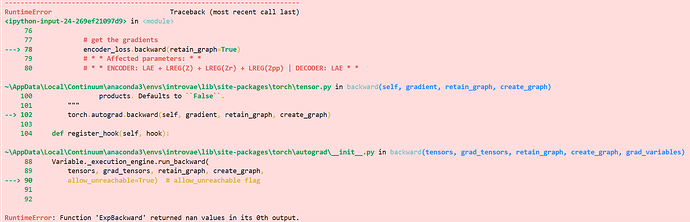
So we are using a residual block for our model, the block looks like this:
class ResidualBlock(nn.Module):
def __init__(self, in_channels, out_channels, triplet=True):
super(ResidualBlock, self).__init__()
# conv layers
self.conv_res = nn.Conv2d(in_channels, out_channels, kernel_size=1, padding=0)
self.conv_1 = nn.Conv2d(in_channels, out_channels, kernel_size=3, padding=1)
self.conv_2 = nn.Conv2d(out_channels, out_channels, kernel_size=3, padding=1)
# initialization
kaiming_normal_(self.conv_res.weight, nonlinearity='relu')
kaiming_normal_(self.conv_1.weight, nonlinearity='relu')
kaiming_normal_(self.conv_2.weight, nonlinearity='relu')
# batch norm layers
self.bn1 = nn.BatchNorm2d(out_channels)
self.bn2 = nn.BatchNorm2d(out_channels)
self.bn3 = nn.BatchNorm2d(out_channels)
# activations
self.relu = nn.ReLU()
# flags
self.triplet = triplet
def forward(self, x):
# out placeholder
out = None
# if the block is a triplet
if self.triplet:
# first stage
out_1 = self.conv_1(x)
out_1 = self.bn1(out_1)
out_1 = self.relu(out_1)
# second stage
out_2 = self.conv_2(out_1)
out_2 = self.bn2(out_2)
# third stage - residual connection
out_res = self.conv_res(x)
# if the block is a twin
else:
# first stage
out_1 = self.conv_1(x)
out_1 = self.bn1(out_1)
out_1 = self.relu(out_1)
# second stage
out_2 = self.conv_2(out_1)
out_2 = self.bn2(out_2)
# add the activations
if self.triplet:
out = out_2 + out_res
else:
out = out_2 + x
# final activation function
out = self.relu(out)
return out
The model is basically a stack of this residual blocks. Here is the encoder part of it:
class Encoder(nn.Module):
def __init__(self):
super(Encoder, self).__init__()
# conv layer
# using 3 by 3 receptive field
self.conv = nn.Conv2d(in_channels=3, out_channels=16, kernel_size=3, stride=1, padding=1)
# initialize the conv layer
kaiming_normal_(self.conv.weight.data, nonlinearity='relu') # init with kaiming he
# residual blocks
self.res1 = ResidualBlock(in_channels=16, out_channels=32)
self.res2 = ResidualBlock(in_channels=32, out_channels=64)
self.res3 = ResidualBlock(in_channels=64, out_channels=128)
self.res4 = ResidualBlock(in_channels=128, out_channels=256)
self.res5 = ResidualBlock(in_channels=256, out_channels=512)
# fully connected layer
self.fc1024 = nn.Linear(in_features=8192, out_features=1024)
# initialize the fc
kaiming_normal_(self.fc1024.weight.data, nonlinearity='relu')
# pool layer
self.pool = nn.AvgPool2d(kernel_size=2)
def forward(self, x):
# x is 3x256x256
# input block
out = self.conv(x) # 16x256x256
out = self.pool(out) # 16x128x128
print("CONV MAX ACT: ", out.max())
# residual block 1
out = self.res1(out) # 32x128x128
out = self.pool(out) # 32x64x64
print("RES1 MAX ACT: ", out.max())
# residual block 2
out = self.res2(out) # 64x64x64
out = self.pool(out) # 64x32x32
print("RES2 MAX ACT: ", out.max())
# residual block 3
out = self.res3(out) # 128x32x32
out = self.pool(out) # 128x16x16
print("RES3 MAX ACT: ", out.max())
# residual block 4
out = self.res4(out) # 256x16x16
out = self.pool(out) # 256x8x8
print("RES4 MAX ACT: ", out.max())
# residual block 5
out = self.res5(out) # 512x8x8
out = self.pool(out) # 512x4x4
print("RES5 MAX ACT: ", out.max())
print("CONV MAX WEIGHT: ", self.conv.weight.max(), self.conv.bias.max())
print("RES1 MAX WEIGHTS: ", self.res1.conv_1.weight.max(), self.res1.conv_2.weight.max(), self.res1.conv_res.weight.max())
print("RES2 MAX WEIGHTS: ", self.res2.conv_1.weight.max(), self.res2.conv_2.weight.max(), self.res2.conv_res.weight.max())
print("RES3 MAX WEIGHTS: ", self.res3.conv_1.weight.max(), self.res3.conv_2.weight.max(), self.res3.conv_res.weight.max())
print("RES4 MAX WEIGHTS: ", self.res4.conv_1.weight.max(), self.res4.conv_2.weight.max(), self.res4.conv_res.weight.max())
print("RES5 MAX WEIGHTS: ", self.res5.conv_1.weight.max(), self.res5.conv_2.weight.max(), self.res5.conv_res.weight.max())
print("OUT_PRE MAX: ", out.max())
# reshape
out = out.view((out.shape[0], -1))
# fully connected
out = self.fc1024(out)
print("OUT_SHAPE: ", out.shape, "OUT: ", out, "OUT MAX: ", out.max())
# get the mean and log-variance
mu = out[:, :512]
logvar = out[:, 512:]
# reparametrization trick
z = self.reparameterize(mu, logvar)
# return the latent vector, mu and logvar
return z, mu, logvar
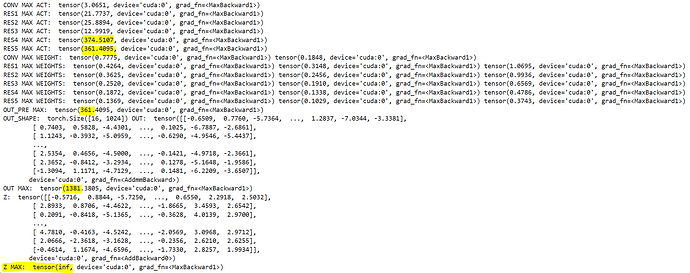
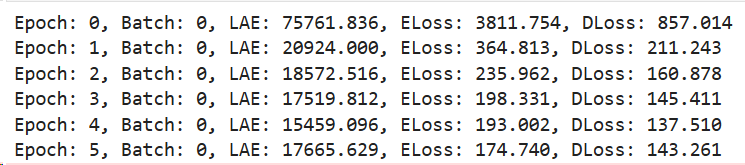
The print statements print the next stats:
This is on the first batch of the first epoch.