I’ve been training a model and have not been getting the results that I expect. I have a suspicion that it might be due to vanishing/exploding gradients, but would like to verify this somehow. How might I go about inspecting the gradients of my model to verify that this is in fact the case?
I am not sure how to identify/verify exploding gradients, you could try gradient clipping using something like below that will prevent the gradients from going aboard:
torch.nn.utils.clip_grad_norm(model.parameters()) as shown in:
https://github.com/pytorch/examples/blob/master/word_language_model/main.py#L138 and see if that makes any difference.
I think the canonical reference for finding bad gradients is this snippet by Adam Paszke:
It checks for NaN (by using x!=x if and only if x is NaN) and very large gradients, but you could easily adapt is_bad_grad to best fit your purpose.
Best regards
Thomas
Thanks, I’ll definitely be taking a look into this. Very helpful.
In my case, I use this simple code to see values of gradient, particularly for rnn:
for p,n in zip(rnn.parameters(),rnn._all_weights[0]):
if n[:6] == 'weight':
print('===========\ngradient:{}\n----------\n{}'.format(n,p.grad))
That’s exactly what I was looking for. Thanks!
This snippet appears not to work with Python 3.7 and PyTorch 1.5.0. Is there a new “canonical” reference?
The new way of getting bad grads is the anomaly mode. But since you asked, I put up a Notebook of Adam’s bad_grad_viz adapted to modern PyTorch at https://github.com/t-vi/pytorch-tvmisc/blob/master/visualize/bad_grad_viz.ipynb.
@jel158 @tom
sorry if I want to see the gradient of my model as follow how I should use the code?
class Generator994(nn.Module):
def __init__(self,ngpu,nz,ngf):
super(Generator994, self).__init__()
self.ngpu=ngpu
self.nz=nz
self.ngf=ngf
self.l1= nn.Sequential(
nn.ConvTranspose2d(self.nz, self.ngf * 8, 3, 1, 0, bias=False),
nn.BatchNorm2d(self.ngf * 8),
nn.ReLU(True),)
self.l2=nn.Sequential(nn.ConvTranspose2d(self.ngf * 8, self.ngf * 4, 3, 1, 0, bias=False),
nn.BatchNorm2d(self.ngf * 4),
nn.ReLU(True),)
self.l3=nn.Sequential(nn.ConvTranspose2d( self.ngf * 4, self.ngf * 2, 3, 1, 0, bias=False),
nn.BatchNorm2d(self.ngf * 2),
nn.Sigmoid(),)
self.l4=nn.Sequential(nn.ConvTranspose2d( self.ngf*2, 1, 3, 1, 0, bias=False),nn.Sigmoid()
)
def forward(self, input):
out=self.l1(input)
out=self.l2(out)
out=self.l3(out)
out=self.l4(out)
return out
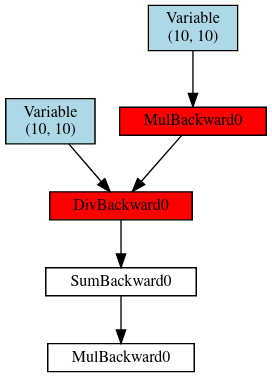
The idea of the linked code is to just run your model as usual to give some function loss.
Then you add the following before and after the backward:
get_dot = register_hooks(loss)
loss.backward()
dot = get_dot()
and then dot contains a dot graph object that you can display in Jupyter or render.
Note that it might get large fast, but for a convnet like you posted it should be OK.
Then the red items are the ones that are problematic. (You could adapt the criterion by changing is_bad_grad.
The other question to ask is, of course, why you’re having exploding gradient issue and what you would do if you knew “where” is happens.
Best regards
Thomas
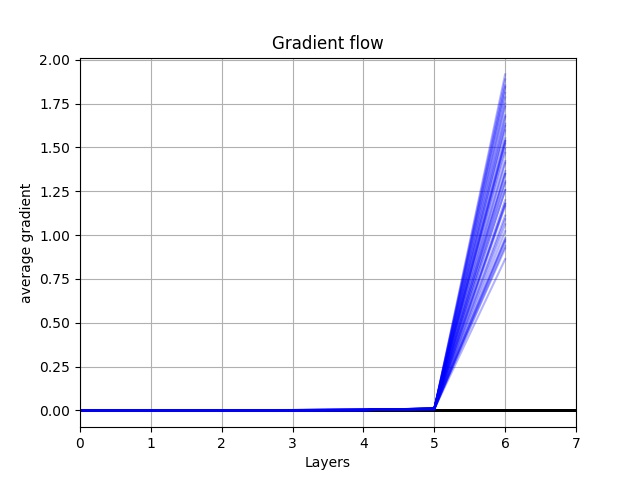
I appreciate your explanation. I get this graph as my gradient flow, do you think the number of layers should be cut? because the gradient in the earliest layer are very small. what is the feature of good gradient flow?
I get error ‘Net’ object has no attribute ‘all_weights’…
_all_weights is an internal attribute of RNNBase and I would not recommend to rely on the usage of these internal objects, as they might change without any deprecation warning.
That being said, what is your exact use case? If you want to filter out some parameters of the RNN you could try to use model.rnn.named_parameters() instead.
Let me know, if this would work for you.
Dear Piotr,
Oh it is clear then - I do not use RNNBase. I was just looking for the way to see what is happening to the gradients in my network to overall have an idea what is going on in it. If there is a quick pointer you could share, I would be grateful.
Best,
Alice
You could manually check all gradients e.g. via:
for name, param in model.named_parameters():
print(name, param.grad.norm())
(or any other stats, if norm is not desired).
However, this approach would be quite limited and more sophisticated algorithms for model interpretability can be applied by e.g. Captum.
Also, you might want to plot the histograms of the gradients in TensorBoard or any other visualization tool, which should give you more insight into the model training.
How am I supposed to interpret it that anomaly mode is pointing to torch.nn.mse_loss() as the place where the gradient goes bad?
Similarly the graphing code posted by @tom above show red nodes from beginning (after input variable) to end (mse loss function).
You could check the input to mse_loss and its gradients (use out.retain_grad() to keep the gradient).
Best regards
Thomas
Thanks!
You are right, it seems like the weights are updated such that an intermediate tensor becomes nan somewhere in the model.
I’ve tried implementing this code roughly as follows:
loss = self.get_loss(x, x_hat)
get_dot = register_hooks(loss)
loss.backward()
dot = get_dot()
I’m able to generate graphs for a few batches, however after about 10 batches I receive an assertion error stemming from the fact that a backward gradient function is not in the function dictionary. When printing the size of the dictionary, I notice that the number of functions in the dictionary remains constant for a number of batches, and then drops suddenly. Do you have any idea why register_grad in hook_cb is not adding the full set of functions to the dictionary?
Thanks!
I would try batch normalization, it may help the gradient flow reach the early layers before dying out.
Regarding what a good gradient flow looks like, recall that the gradient influences how much the model is able to learn from an instance of data. Thus, a healthy gradient flow should be non-zero (mostly) from the top layer all the way to the input layer. Otherwise, the weights in the earlier layers will not update at all.
I find it more intuitive to examine the graph from right to left since this is actually how one would compute gradients during backpropagation.