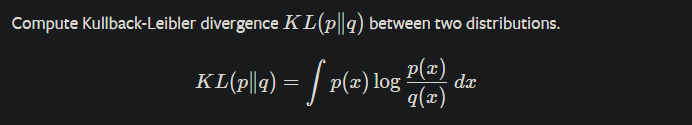- return apply_loss_reduction(output, reduction);
- }
- Tensor margin_ranking_loss(const Tensor& input1, const Tensor& input2, const Tensor& target, double margin, int64_t reduction) {
- auto output = (-target * (input1 - input2) + margin).clamp_min_(0);
- return apply_loss_reduction(output, reduction);
- }
- Tensor kl_div(const Tensor& input, const Tensor& target, int64_t reduction) {
- auto zeros = at::zeros_like(target);
- auto output_pos = target * (at::log(target) - input);
- auto output = at::where(target > 0, output_pos, zeros);
- return apply_loss_reduction(output, reduction);
- }
- Tensor kl_div_backward_cpu(const Tensor& grad, const Tensor& input, const Tensor& target, int64_t reduction) {
- auto grad_input = at::zeros_like(input);
- auto grad_expand = grad.expand_as(input);
- AT_DISPATCH_FLOATING_TYPES(input.scalar_type(), "kl_div_backward_cpu", [&]() {
- at::CPU_tensor_apply3<scalar_t, scalar_t, scalar_t>(
- grad_input,