Hi @tom, it’s really cool that you’re getting interested in this problem. I just want to avoid you going down blind alley’s.
The paper "Stochastic Optimization for Large-scale Optimal Transport " https://arxiv.org/abs/1605.08527, is a conference paper, and they’re usually a bit of a wild card. I spent a while on the code, audeg/StochasticOT, and it’s probably more suited to information retrieval rather than actually training a network like @smth’s GAN code, or as a layer in one. So I think I made a mistake, and it’s perhaps not such a good idea implementing it as a layer.
I don’t want to mislead you - it’s probably a good idea to work on something that’s been proven to be useful. If you simply try to reproduce Chiyuan Zhang (pluskid) Wassertein.jl layer, in the code at the top of this thread, that would be a safe thing to do. That’s something that were reasonably sure work’s, and if you get it working in PyTorch it would be something that you could reference to, and reuse in the future.
Anyway, starting with the easy stuff - at the moment I can’t get the entropy regularised version in,
https://github.com/rflamary/POT/blob/master/examples/Demo_1D_OT.ipynb.
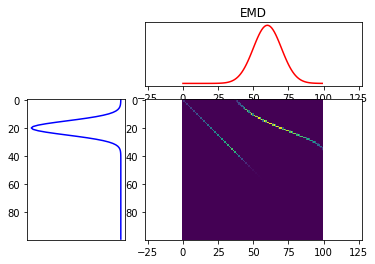
and the exact EMD solver used in PyEMD to give roughly the same number’s? What I mean is, (I think) the functions in the POT library return the transportation plan matrix T, and to get the actual wasserstein/EMD divergence, you calculate, the inner product,
<T,M> = trace(T'M)
Where M is the ground metric matrix. But I get different numbers from both libraries, so I’m not sure which is right? I know you need to tune the regularization parameter lambda, but it should be easy to do that using downhill simplex/Nelder Mead
I want to get these test cases to reconcile - that way we’ve got something to check against when we try to code things, and do more complicated stuff in PyTorch. Otherwise, it’s too easy to make a mistake, without something solid to test against. 

 , I think I’m starting to understand what’s going on!
, I think I’m starting to understand what’s going on!