I use a simple trick. I record the average gradients per layer in every training iteration and then plotting them at the end. If the average gradients are zero in the initial layers of the network then probably your network is too deep for the gradient to flow.
So this is how I do it -
- This API to plot
def plot_grad_flow(named_parameters):
ave_grads = []
layers = []
for n, p in named_parameters:
if(p.requires_grad) and ("bias" not in n):
layers.append(n)
ave_grads.append(p.grad.abs().mean())
plt.plot(ave_grads, alpha=0.3, color="b")
plt.hlines(0, 0, len(ave_grads)+1, linewidth=1, color="k" )
plt.xticks(range(0,len(ave_grads), 1), layers, rotation="vertical")
plt.xlim(xmin=0, xmax=len(ave_grads))
plt.xlabel("Layers")
plt.ylabel("average gradient")
plt.title("Gradient flow")
plt.grid(True)
- plug this API after the loss.backward() during the training as follows -
loss = self.criterion(outputs, labels)
loss.backward()
plot_grad_flow(model.named_parameters())
- Results
bad gradient flow -
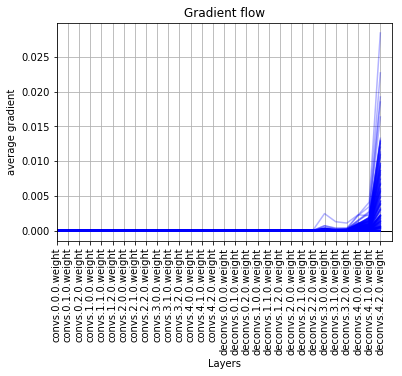
kinda good gradient flow -
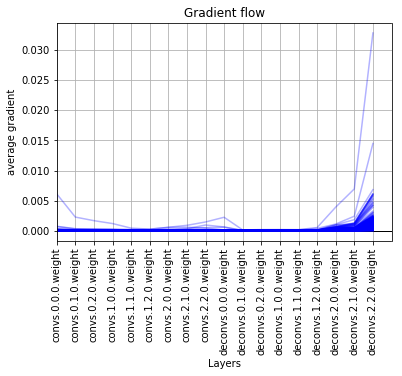
good gradient flow -
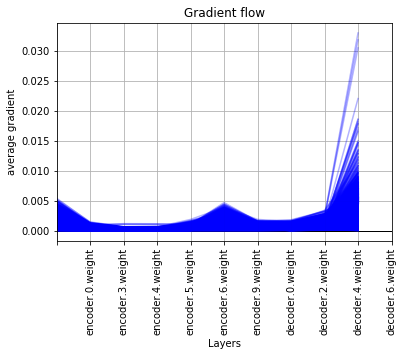
PS : This may not be the best way though…